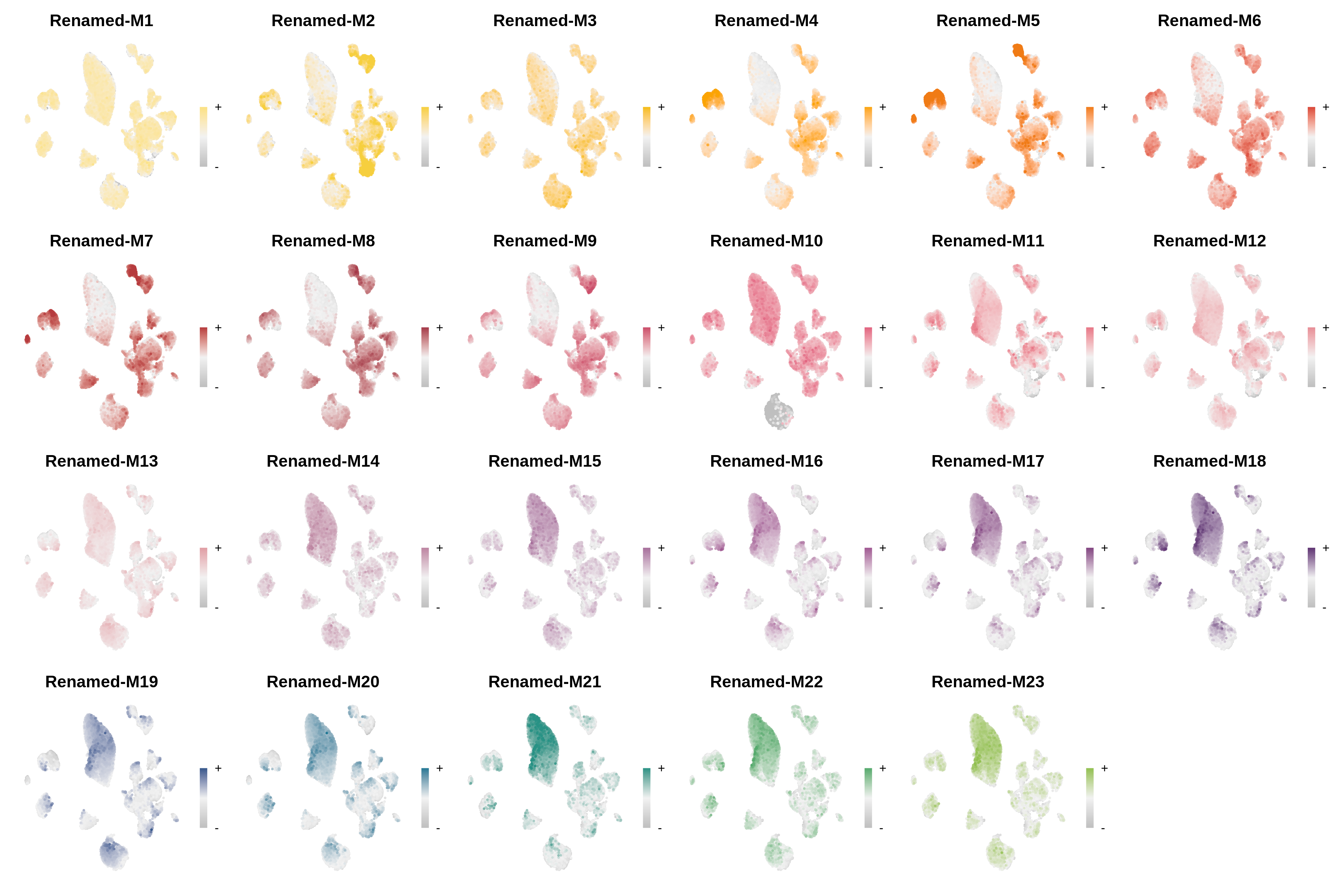
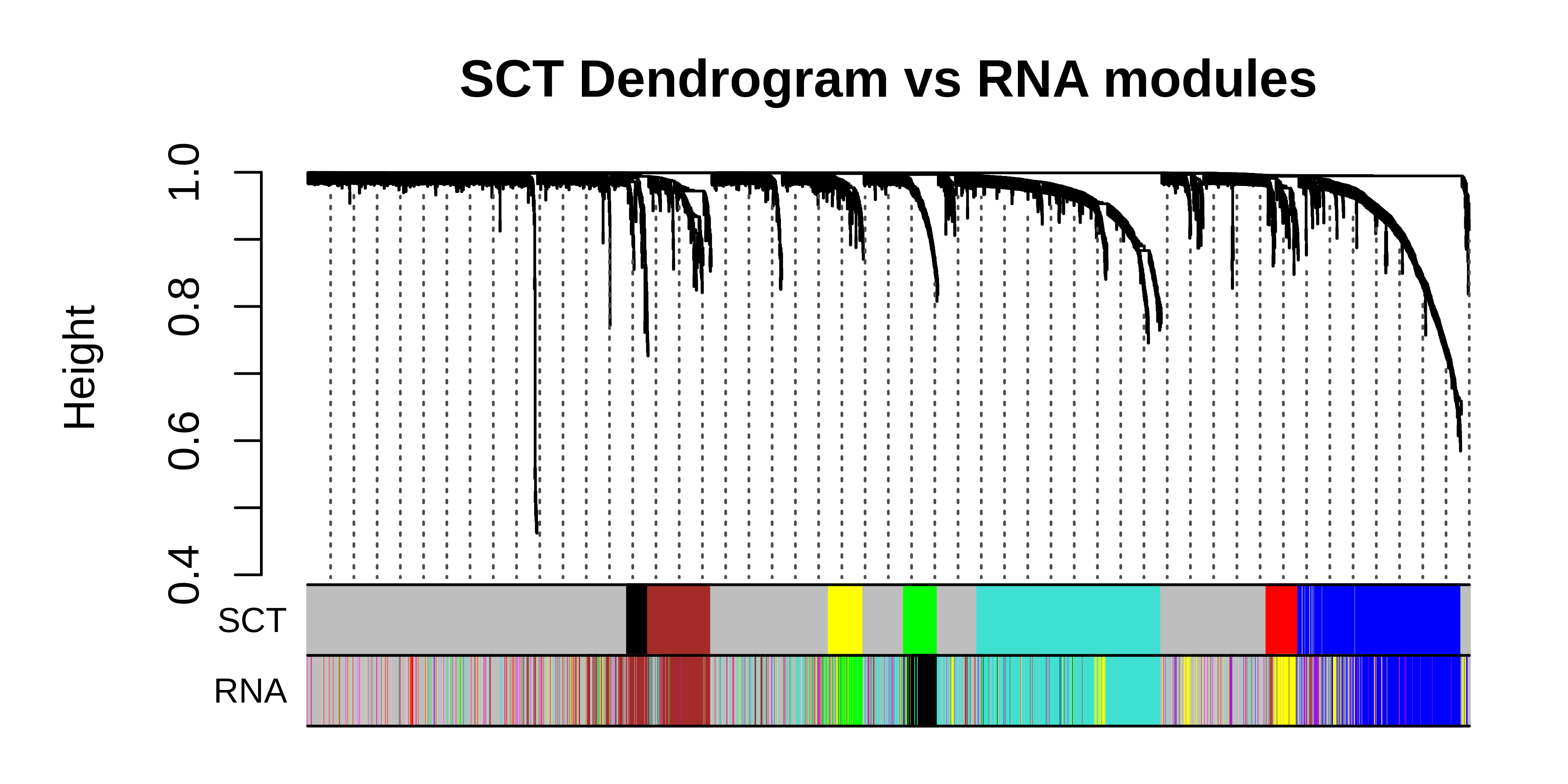
hdWGCNA Tutorials
Welcome to the hdWGCNA documentation! This page provides an overview of all available tutorials organized by topic.
Getting Started
Introduction to hdWGCNA
Learn about the background, motivation, and key concepts behind hdWGCNA.
Quick Start Guide
Get up and running with hdWGCNA in under 10 minutes with a minimal working example.
Algorithm and Mathematical Framework
Detailed explanation of the algorithms and mathematical foundations underlying hdWGCNA, including metacell aggregation, soft power thresholding, and module detection.
Core Tutorials
hdWGCNA in Single-Cell Data
Comprehensive tutorial covering co-expression network analysis in scRNA-seq data.
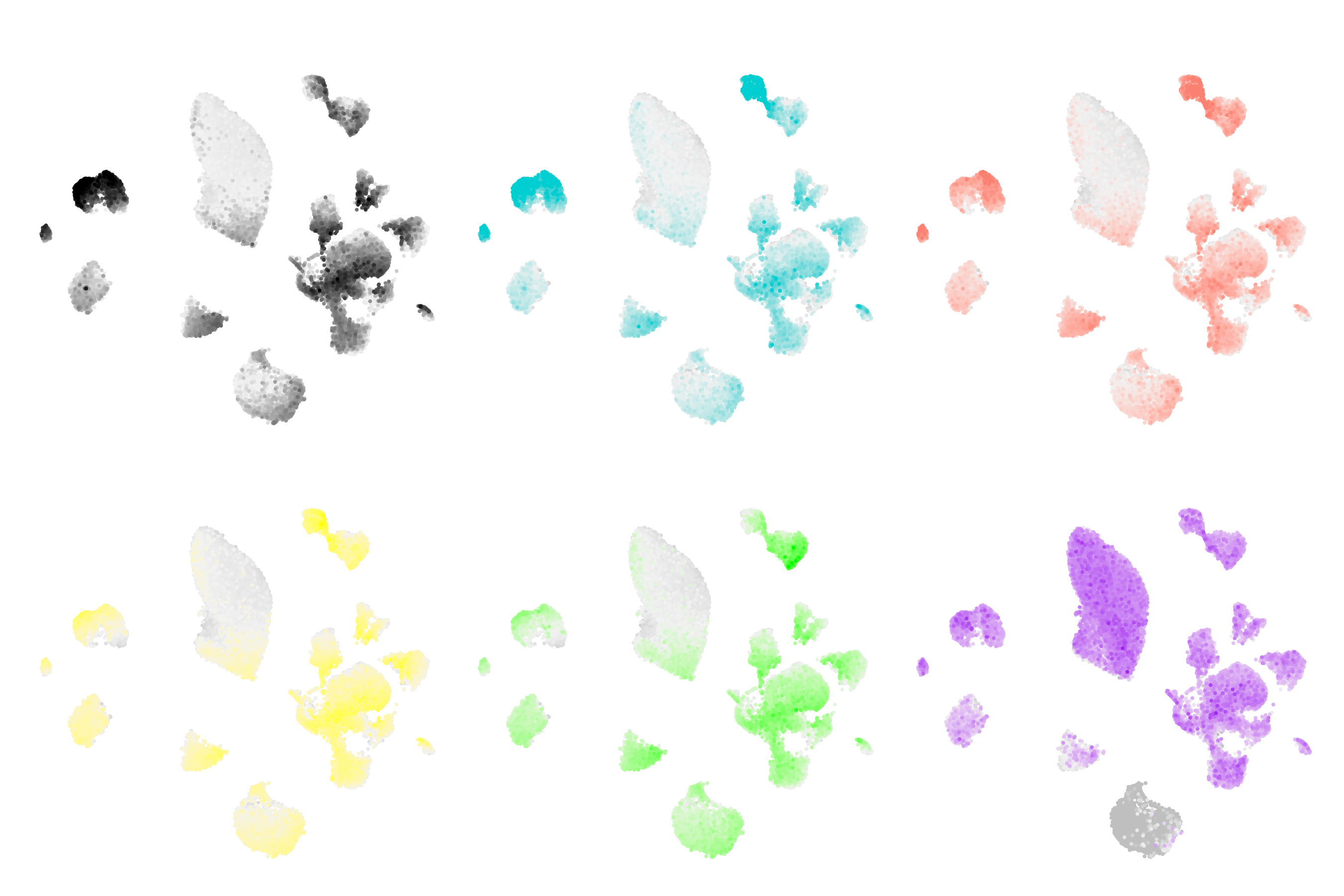
hdWGCNA in Spatial Transcriptomics
Learn how to construct co-expression networks from spatial transcriptomics data.

Biological Context
Differential Module Eigengene (DME) Analysis
Compare module eigengenes between experimental groups to identify condition-specific modules.
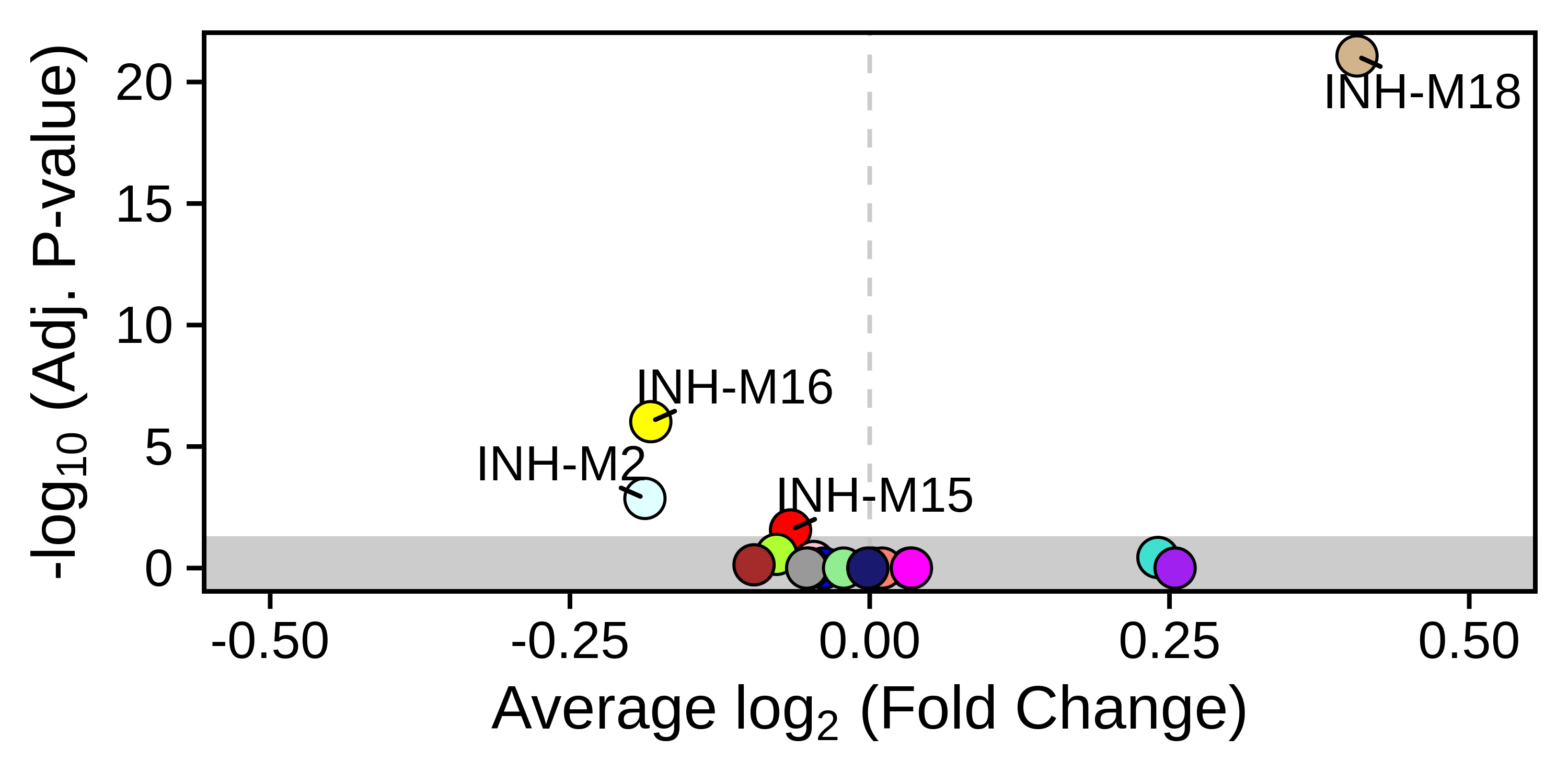
Cross-Dataset Analysis
Projecting Modules to New Datasets
Project co-expression modules from a reference to a query dataset.
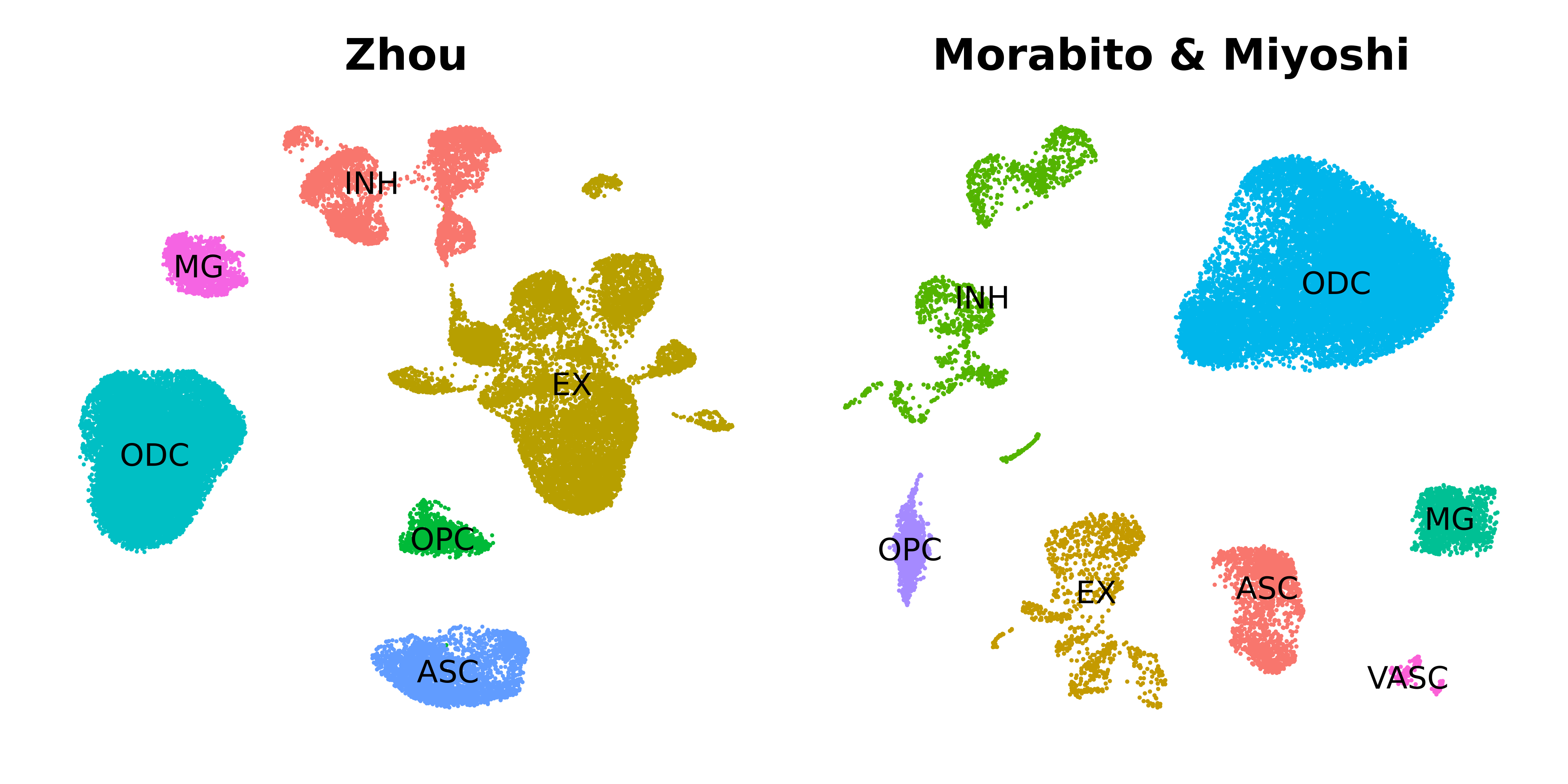
Cross-Species and Cross-Modality Analysis
Special cases for projection when species or data modality differ.
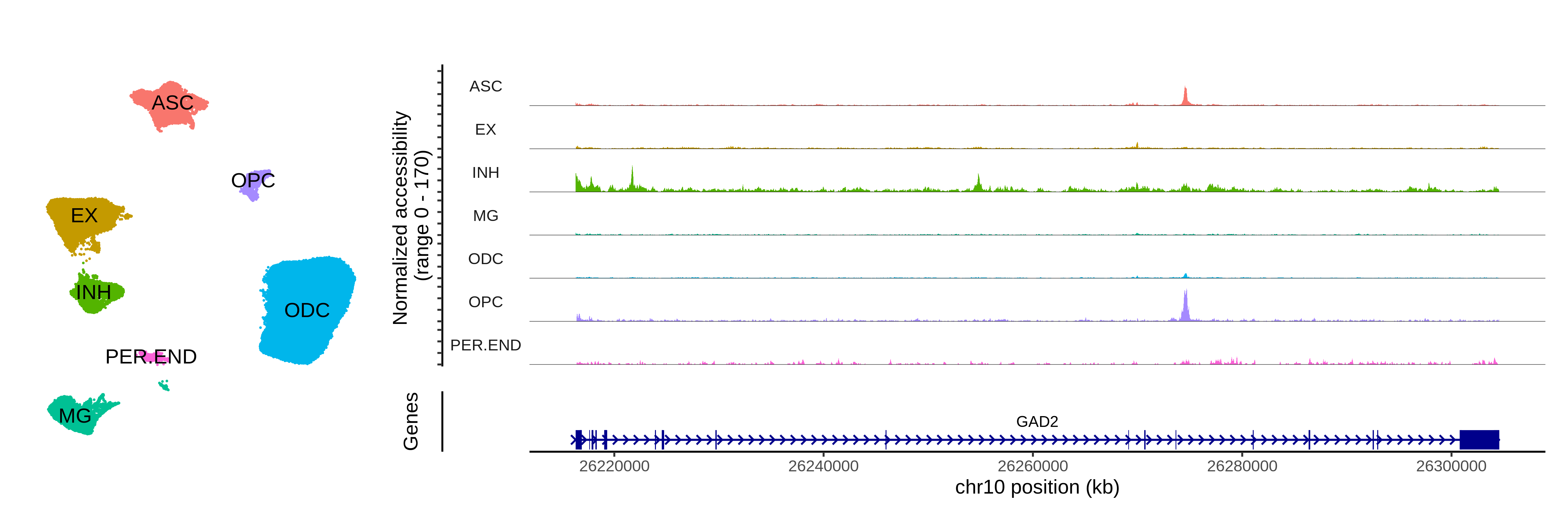
Advanced Topics
Transcription Factor Network Analysis
Infer TF regulatory networks using XGBoost-based modeling.
PPI Network Integration
Integrate co-expression networks with protein-protein interaction data.
Consensus Network Analysis
Combine multiple datasets for robust network construction.
Pseudobulk Analysis
Use hdWGCNA with pseudobulk expression profiles.
Pseudotime Dynamics
Study module dynamics along pseudotime trajectories.
Need Help?
- GitHub Issues: Report bugs or request features
- Function Reference: Complete function documentation