Algorithm Details and Mathematical Framework
Zaoqu Liu
2026-01-23
Source:vignettes/algorithm-details.Rmd
algorithm-details.RmdMathematical Framework
This vignette provides a detailed description of the mathematical foundations underlying the MultiK algorithm.
1. Consensus Clustering
1.1 Subsampling Strategy
Given a dataset with cells, MultiK performs subsampling iterations. In each iteration :
- Randomly sample cells without replacement (default )
- Apply the standard Seurat clustering pipeline across resolution parameters
1.2 Consensus Matrix Construction
For each cluster number , we construct a consensus matrix :
where is the number of runs that yielded exactly clusters.
Properties of the consensus matrix:
- (a cell always co-clusters with itself)
- (symmetric)
- implies cells and always cluster together
- implies cells never cluster together
1.3 Visualizing the Consensus Matrix
library(ggplot2)
# Simulate a consensus matrix with 3 clear clusters
set.seed(42)
n <- 60
blocks <- c(rep(1, 20), rep(2, 20), rep(3, 20))
mat <- matrix(0, n, n)
for (i in 1:n) {
for (j in 1:n) {
if (blocks[i] == blocks[j]) {
mat[i, j] <- runif(1, 0.85, 1.0)
} else {
mat[i, j] <- runif(1, 0, 0.15)
}
}
}
diag(mat) <- 1
mat[lower.tri(mat)] <- t(mat)[lower.tri(mat)]
# Reorder by cluster
ord <- order(blocks)
mat_ord <- mat[ord, ord]
df_cons <- expand.grid(Cell1 = 1:n, Cell2 = 1:n)
df_cons$value <- as.vector(mat_ord)
ggplot(df_cons, aes(x = Cell1, y = Cell2, fill = value)) +
geom_tile() +
scale_fill_gradient2(low = "#3498DB", mid = "white", high = "#E74C3C",
midpoint = 0.5, limits = c(0, 1),
name = "Co-clustering\nProbability") +
labs(x = "Cell Index", y = "Cell Index",
title = "Consensus Matrix Heatmap",
subtitle = "Clear block structure indicates stable clusters") +
coord_fixed() +
theme_minimal(base_size = 12) +
theme(plot.title = element_text(face = "bold"),
panel.grid = element_blank())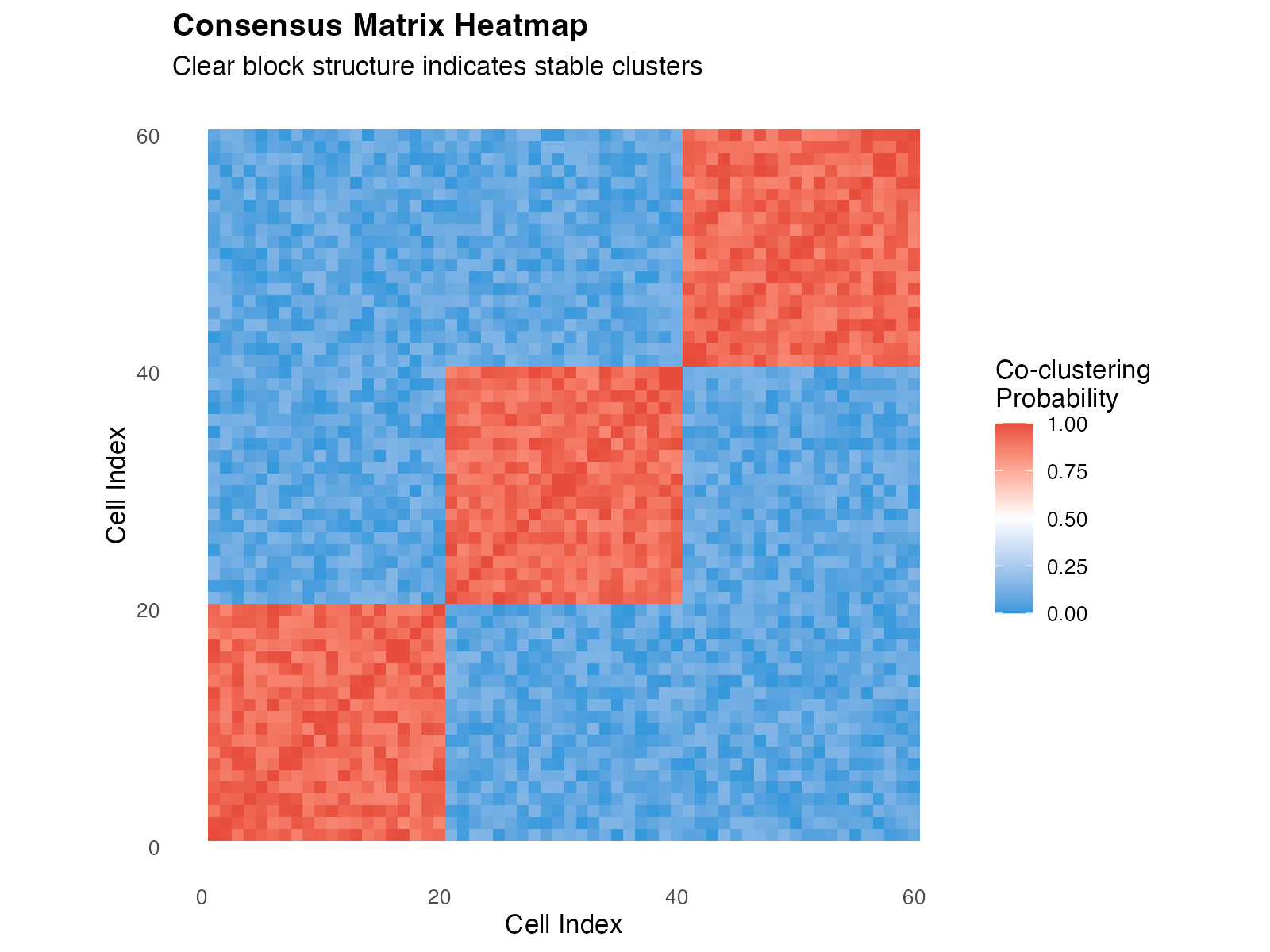
Example consensus matrix showing clear block-diagonal structure indicating well-separated clusters.
2. Proportion of Ambiguous Clustering (PAC)
2.1 Definition
The PAC metric quantifies clustering ambiguity by measuring the proportion of cell pairs with intermediate consensus values:
where and are default thresholds.
2.2 Interpretation
| PAC Value | Interpretation |
|---|---|
| PAC ≈ 0 | Clear cluster structure, cells consistently assigned |
| PAC ≈ 1 | Ambiguous clustering, unstable cell assignments |
2.3 PAC Visualization
# Simulate PAC values for different K
df_pac <- data.frame(
K = factor(2:8),
PAC = c(0.12, 0.05, 0.08, 0.18, 0.35, 0.52, 0.68)
)
df_pac$rPAC <- df_pac$PAC / max(df_pac$PAC)
p1 <- ggplot(df_pac, aes(x = K, y = PAC)) +
geom_col(fill = "#3498DB", color = "black", width = 0.7) +
geom_hline(yintercept = 0.1, linetype = "dashed", color = "red") +
labs(x = "Number of Clusters (K)", y = "PAC",
title = "Absolute PAC") +
theme_bw(base_size = 12) +
theme(plot.title = element_text(face = "bold"))
p2 <- ggplot(df_pac, aes(x = K, y = rPAC)) +
geom_col(fill = "#E74C3C", color = "black", width = 0.7) +
labs(x = "Number of Clusters (K)", y = "Relative PAC (rPAC)",
title = "Relative PAC (normalized)") +
theme_bw(base_size = 12) +
theme(plot.title = element_text(face = "bold"))
cowplot::plot_grid(p1, p2, ncol = 2)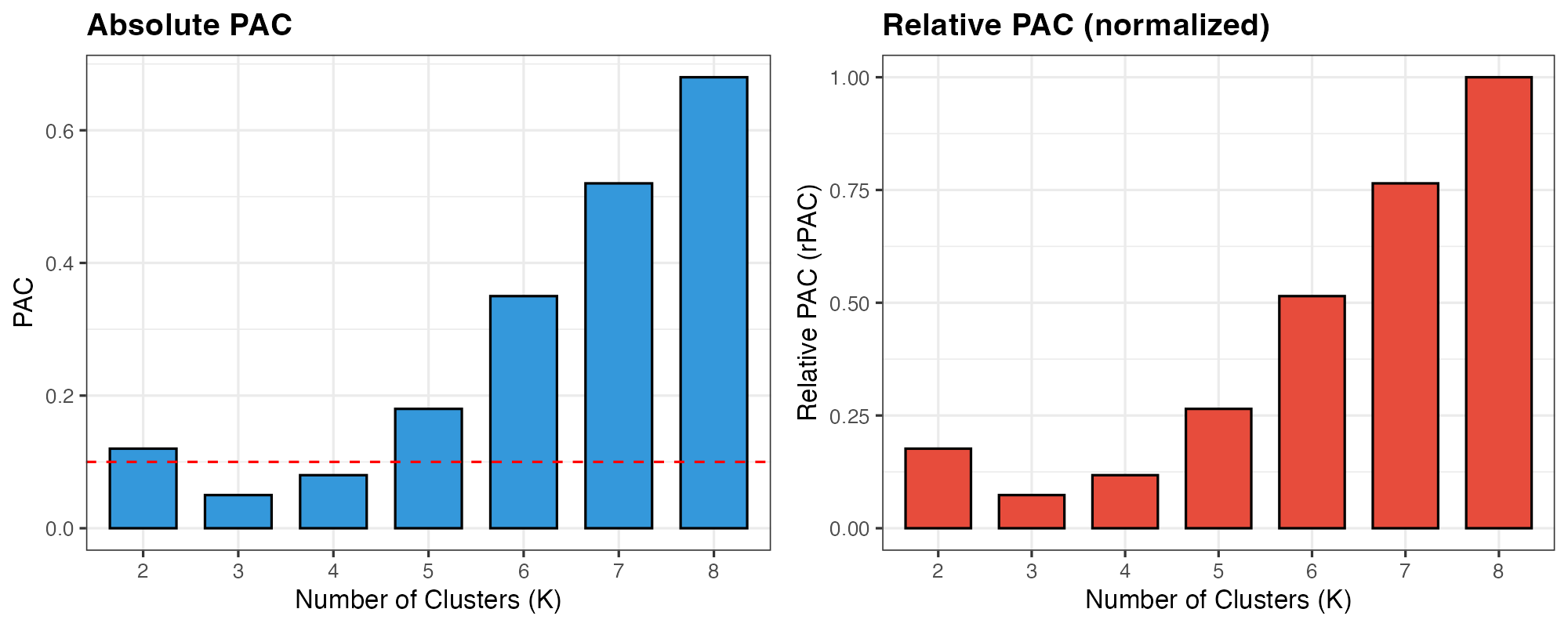
PAC values across different K. Lower PAC indicates more stable clustering.
3. Optimal K Selection
3.1 Multi-Objective Optimization
Optimal K should satisfy two criteria:
- High frequency: K appears consistently across resolution parameters
- Low rPAC: Clustering is stable (low ambiguity)
3.2 Pareto Frontier
We identify optimal K values using Pareto optimality. A point is Pareto-optimal if no other point satisfies:
where: - (stability) -
set.seed(42)
df <- data.frame(
K = factor(2:8),
stability = c(0.88, 0.95, 0.92, 0.82, 0.65, 0.48, 0.32),
frequency = c(25, 120, 85, 95, 110, 75, 40)
)
# Find Pareto optimal points
pareto <- c()
for (i in 1:nrow(df)) {
dominated <- FALSE
for (j in 1:nrow(df)) {
if (i != j) {
if (df$stability[j] >= df$stability[i] && df$frequency[j] >= df$frequency[i] &&
(df$stability[j] > df$stability[i] || df$frequency[j] > df$frequency[i])) {
dominated <- TRUE
break
}
}
}
if (!dominated) pareto <- c(pareto, i)
}
df$optimal <- 1:7 %in% pareto
ggplot(df, aes(x = stability, y = frequency)) +
geom_point(aes(color = optimal), size = 5) +
geom_text(aes(label = paste0("K=", K)), vjust = -1.2, size = 4) +
scale_color_manual(values = c("FALSE" = "gray50", "TRUE" = "#E74C3C"),
guide = "none") +
geom_vline(xintercept = 0.9, linetype = "dashed", alpha = 0.5) +
geom_hline(yintercept = 100, linetype = "dashed", alpha = 0.5) +
annotate("rect", xmin = 0.9, xmax = 1, ymin = 100, ymax = 130,
fill = "green", alpha = 0.1) +
annotate("text", x = 0.95, y = 125, label = "Ideal\nRegion", size = 3) +
labs(x = "Stability (1 - rPAC)", y = "Frequency",
title = "Pareto Frontier for Optimal K Selection",
subtitle = "Red points = Pareto-optimal; Green region = ideal") +
xlim(0.25, 1) +
theme_bw(base_size = 12) +
theme(plot.title = element_text(face = "bold"))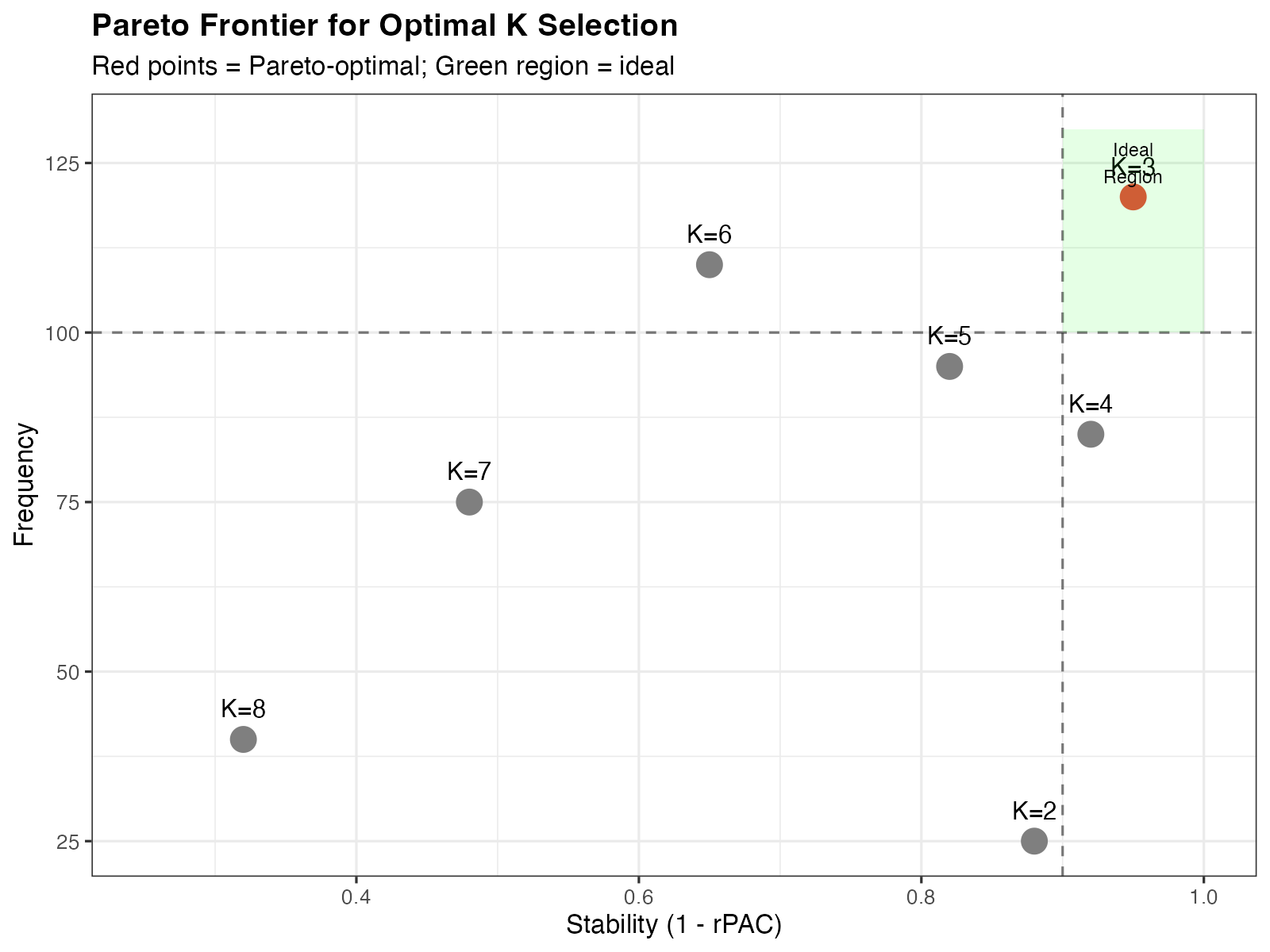
Pareto frontier for K selection. Red points indicate Pareto-optimal K values.
4. SigClust Statistical Testing
4.1 Hypothesis Framework
For each pair of clusters and , SigClust tests:
- : Data come from a single Gaussian distribution
- : Data come from two distinct distributions
4.2 Cluster Index
The test statistic is the cluster index (CI):
where: - are cluster sizes - are cluster centroids - is the pooled covariance estimate
4.3 P-value Heatmap Interpretation
# Create example p-value matrix
pval_mat <- matrix(c(NA, 0.001, 0.015, 0.042,
0.001, NA, 0.003, 0.089,
0.015, 0.003, NA, 0.008,
0.042, 0.089, 0.008, NA), 4, 4, byrow = TRUE)
rownames(pval_mat) <- colnames(pval_mat) <- paste0("Cluster", 1:4)
df_heat <- expand.grid(Cluster1 = rownames(pval_mat),
Cluster2 = colnames(pval_mat))
df_heat$pvalue <- as.vector(pval_mat)
ggplot(df_heat, aes(x = Cluster1, y = Cluster2, fill = pvalue)) +
geom_tile(color = "white", linewidth = 1) +
geom_text(aes(label = ifelse(is.na(pvalue), "", sprintf("%.3f", pvalue))),
size = 5, fontface = "bold") +
scale_fill_gradient2(low = "#27AE60", mid = "#F1C40F", high = "#E74C3C",
midpoint = 0.05, limits = c(0, 0.1),
na.value = "grey90",
name = "P-value") +
labs(x = "", y = "",
title = "Pairwise SigClust P-values",
subtitle = "Green (p < 0.05) = significant separation") +
coord_fixed() +
theme_minimal(base_size = 12) +
theme(plot.title = element_text(face = "bold"),
panel.grid = element_blank(),
axis.text = element_text(size = 11, face = "bold"))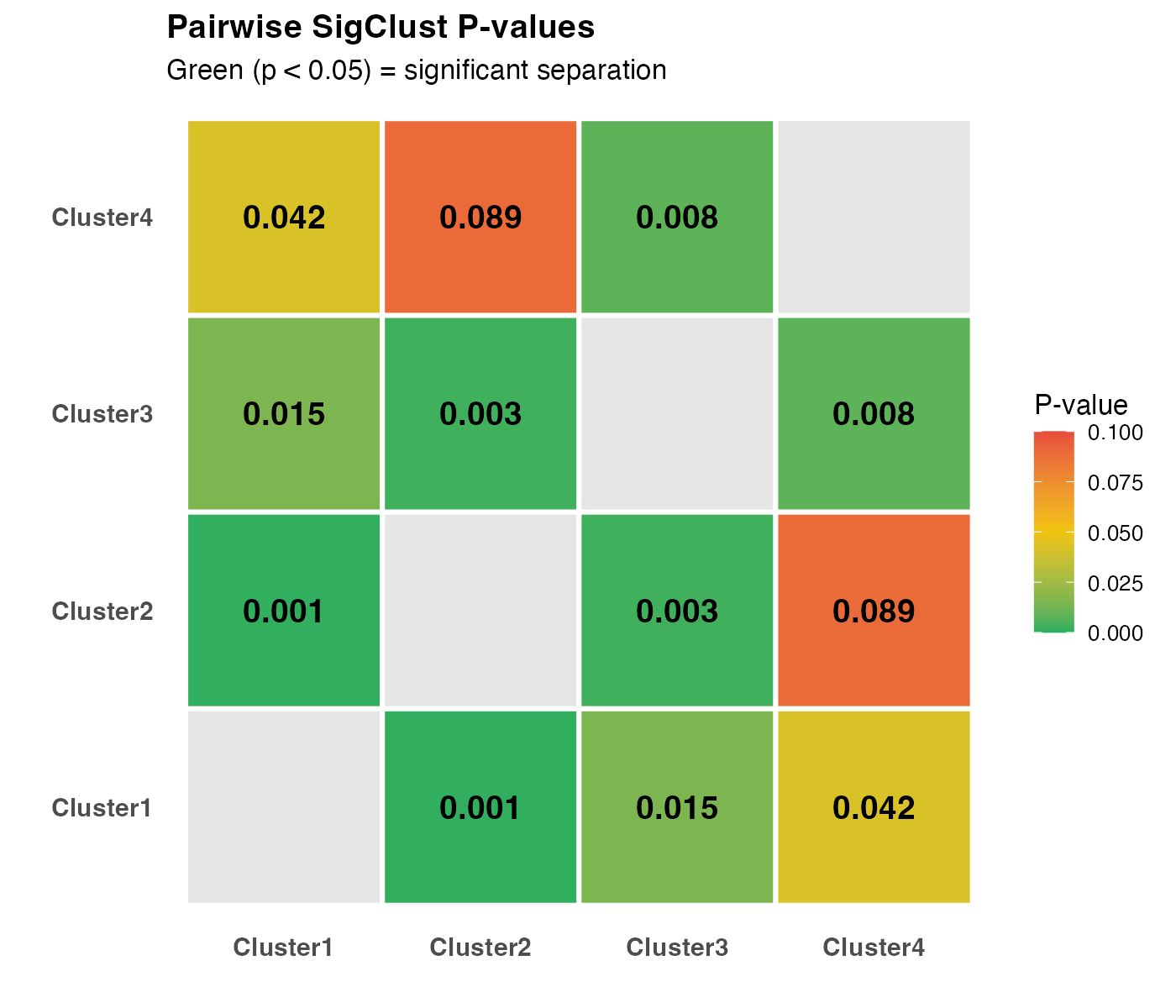
Pairwise SigClust p-value heatmap. Green indicates significant separation.
5. Complete Workflow Visualization
library(ggplot2)
# Create workflow diagram
steps <- data.frame(
x = 1:5,
y = rep(0, 5),
label = c("1. Input\nSeurat Object",
"2. Subsampling\n& Clustering",
"3. Consensus\nMatrix",
"4. PAC &\nOptimal K",
"5. SigClust\nValidation"),
color = c("#3498DB", "#9B59B6", "#E74C3C", "#F1C40F", "#27AE60")
)
ggplot(steps, aes(x = x, y = y)) +
geom_segment(aes(x = 1.3, xend = 4.7, y = 0, yend = 0),
arrow = arrow(length = unit(0.3, "cm"), type = "closed"),
color = "gray40", linewidth = 1.5) +
geom_point(aes(fill = color), shape = 21, size = 20, color = "black", stroke = 2) +
geom_text(aes(label = label), y = -0.15, size = 3.5, lineheight = 0.9) +
scale_fill_identity() +
coord_cartesian(ylim = c(-0.3, 0.15), xlim = c(0.5, 5.5)) +
theme_void() +
labs(title = "MultiK Analysis Workflow") +
theme(plot.title = element_text(hjust = 0.5, face = "bold", size = 14))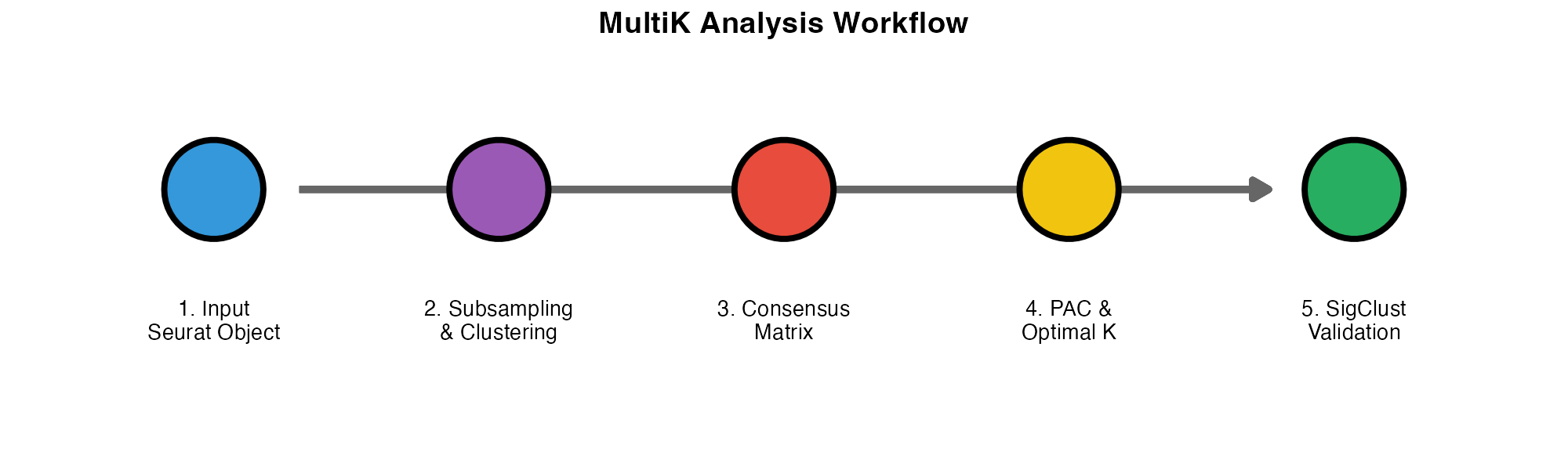
MultiK analysis workflow.
6. Computational Complexity
| Component | Complexity |
|---|---|
| Single clustering run | |
| All subsampling iterations | |
| Consensus matrix | |
| PAC calculation | |
| SigClust (per pair) |
where = cells, = reps, = resolutions, = simulations.
7. Parameter Recommendations
| Parameter | Recommended | Rationale |
|---|---|---|
reps |
100-200 | Sufficient for stable consensus |
pSample |
0.8 | Balance between variation and representation |
resolution |
0.05-2.0 | Covers typical cluster range |
nsim |
100-500 | Accurate p-value estimation |
References
Consensus Clustering: Monti S, et al. (2003) Consensus Clustering. Machine Learning 52:91-118.
PAC Metric: Șenbabaoğlu Y, et al. (2014) Critical limitations of consensus clustering. Scientific Reports 4:6207.
SigClust: Liu Y, et al. (2008) Statistical significance of clustering. JASA 103:1281-1293.
Author
Zaoqu Liu, PhD
- Email: liuzaoqu@163.com
- ORCID: 0000-0002-0452-742X
This vignette is part of the MultiK package. For questions or issues, please visit GitHub.