Performance Optimization Guide
Zaoqu Liu
2026-01-26
Source:vignettes/performance-guide.Rmd
performance-guide.RmdIntroduction
This guide provides strategies for optimizing scVeloR performance when working with large single-cell datasets. We cover memory management, parallel computing, and algorithmic optimizations.
Performance Architecture
scVeloR uses a hybrid architecture for optimal performance:
┌─────────────────────────────────────────────────────────────┐
│ scVeloR Performance Stack │
├─────────────────────────────────────────────────────────────┤
│ R Interface Layer │
│ └── Vectorized R operations (Matrix package) │
│ └── Parallel processing (future/parallel) │
├─────────────────────────────────────────────────────────────┤
│ C++ Core (Rcpp/RcppArmadillo) │
│ └── Cosine similarity computation │
│ └── EM algorithm core │
│ └── KNN computations │
├─────────────────────────────────────────────────────────────┤
│ Sparse Matrix Support │
│ └── Memory-efficient storage │
│ └── Optimized linear algebra │
└─────────────────────────────────────────────────────────────┘Memory Optimization
Sparse Matrix Usage
scVeloR automatically uses sparse matrices when beneficial:
library(Matrix)
# Check sparsity of your data
sparsity <- sum(seurat_obj@assays$RNA@counts == 0) /
length(seurat_obj@assays$RNA@counts)
message(sprintf("Data sparsity: %.1f%%", sparsity * 100))
# Force sparse representation
seurat_obj@assays$RNA@counts <- as(seurat_obj@assays$RNA@counts, "dgCMatrix")Memory Profiling
# Check memory usage
format(object.size(seurat_obj), units = "GB")
# Monitor during analysis
gc() # Garbage collection
memory.size() # Current memory usage (Windows)
pryr::mem_used() # Cross-platformChunked Processing
For very large datasets, process in chunks:
# Split cells into chunks
n_cells <- ncol(seurat_obj)
chunk_size <- 10000
n_chunks <- ceiling(n_cells / chunk_size)
# Process each chunk
results <- list()
for (i in seq_len(n_chunks)) {
start_idx <- (i - 1) * chunk_size + 1
end_idx <- min(i * chunk_size, n_cells)
chunk_obj <- seurat_obj[, start_idx:end_idx]
results[[i]] <- process_velocity_chunk(chunk_obj)
gc() # Clean up after each chunk
}
# Merge results
final_results <- merge_velocity_results(results)Parallel Computing
Using the future Package
library(future)
library(future.apply)
# Check available cores
availableCores()
# Setup parallel backend
plan(multisession, workers = 4) # 4 parallel workers
# Run velocity analysis
seurat_obj <- run_velocity(seurat_obj,
mode = "dynamical",
n_cores = 4)
# Reset to sequential
plan(sequential)Platform-Specific Configuration
# Detect OS and configure
if (.Platform$OS.type == "unix") {
# Unix/Mac: use multicore for shared memory
plan(multicore, workers = availableCores() - 1)
} else {
# Windows: use multisession (separate R processes)
plan(multisession, workers = availableCores() - 1)
}Parallel Best Practices
| Scenario | Recommended Setup |
|---|---|
| < 10K cells | Sequential (overhead > benefit) |
| 10K - 50K cells | 4 workers |
| 50K - 100K cells | 8 workers |
| > 100K cells | Max available - 1 |
# Automatic configuration based on dataset size
n_cells <- ncol(seurat_obj)
if (n_cells < 10000) {
n_workers <- 1
} else if (n_cells < 50000) {
n_workers <- min(4, availableCores() - 1)
} else {
n_workers <- availableCores() - 1
}
if (n_workers > 1) {
plan(multisession, workers = n_workers)
}Algorithmic Optimizations
Gene Selection
Reducing the number of genes dramatically speeds up computation:
# Use fewer genes for speed
seurat_obj <- velocity(seurat_obj,
mode = "dynamical",
n_top_genes = 1000) # Default: 2000
# Or use highly variable genes
hvg <- Seurat::VariableFeatures(seurat_obj)
seurat_obj <- velocity(seurat_obj,
mode = "dynamical",
genes = hvg[1:500]) # Top 500 HVGsNeighbor Graph Approximation
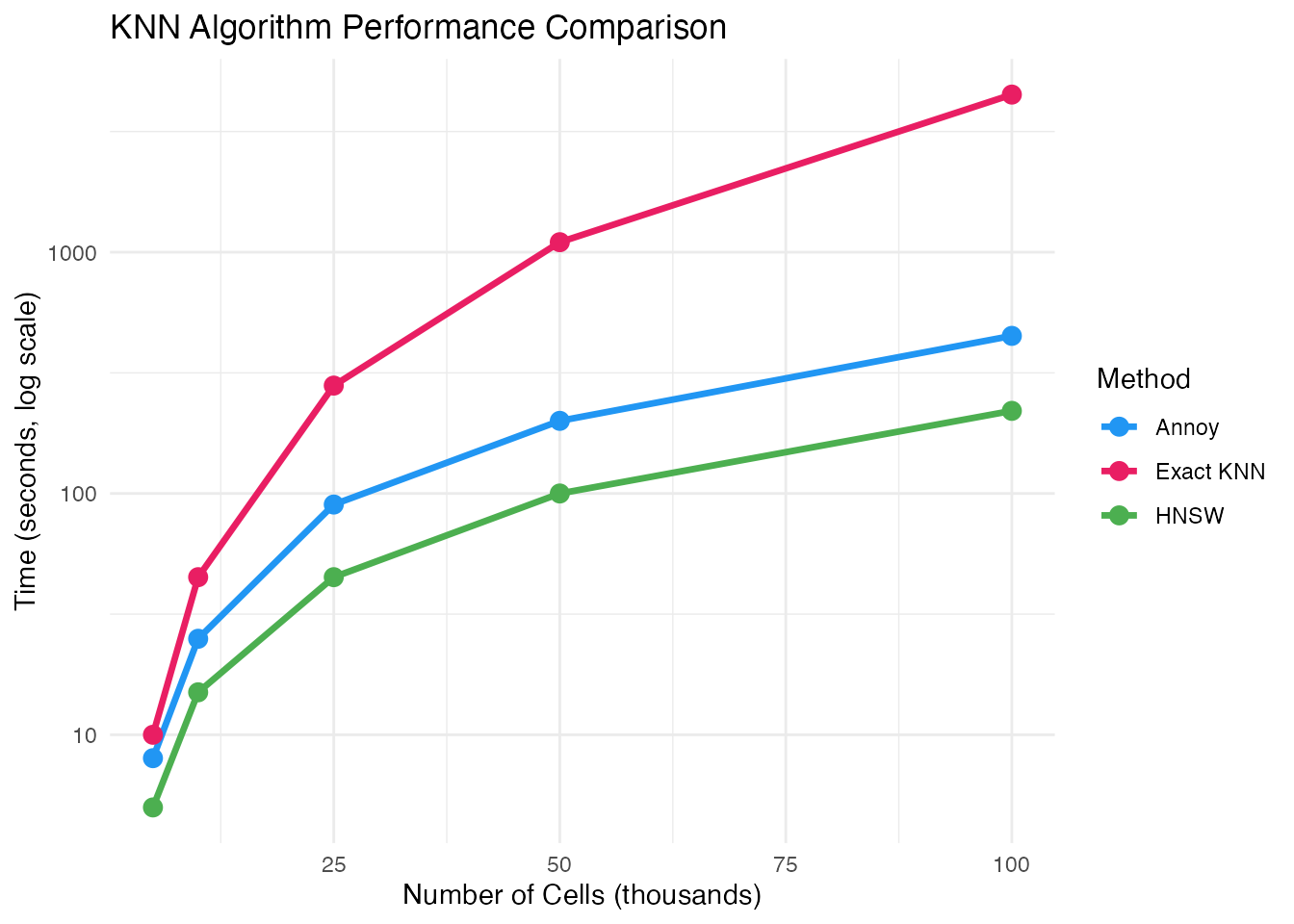
For large datasets, use approximate nearest neighbor algorithms:
# Exact KNN (default, slower for large data)
seurat_obj <- compute_neighbors(seurat_obj,
n_neighbors = 30,
method = "exact")
# Approximate KNN with Annoy (faster)
seurat_obj <- compute_neighbors(seurat_obj,
n_neighbors = 30,
method = "annoy",
n_trees = 50)
# Approximate KNN with HNSW (fastest for very large data)
seurat_obj <- compute_neighbors(seurat_obj,
n_neighbors = 30,
method = "hnsw",
M = 16, ef = 200)EM Algorithm Optimization
# Fewer iterations for faster (less accurate) results
seurat_obj <- recover_dynamics(seurat_obj, max_iter = 5)
# More iterations for better accuracy
seurat_obj <- recover_dynamics(seurat_obj, max_iter = 20)
# Early stopping based on convergence
seurat_obj <- recover_dynamics(seurat_obj,
max_iter = 20,
tol = 1e-4) # Stop if change < tolHardware Recommendations
Profiling Your Analysis
Identifying Bottlenecks
# Use Rprof for detailed profiling
Rprof("velocity_profile.out")
seurat_obj <- velocity(seurat_obj, mode = "dynamical")
Rprof(NULL)
# Analyze results
summaryRprof("velocity_profile.out")Practical Workflow for Large Data
Optimized Pipeline
library(scVeloR)
library(future)
# 1. Configure parallel backend
n_cores <- min(8, availableCores() - 1)
plan(multisession, workers = n_cores)
# 2. Use sparse matrices
seurat_obj@assays$RNA@counts <- as(
seurat_obj@assays$RNA@counts, "dgCMatrix"
)
# 3. Preprocessing with filtering
seurat_obj <- prepare_velocity(
seurat_obj,
min_counts = 30, # Stricter filtering
min_cells = 50,
n_neighbors = 30
)
# 4. Use approximate KNN
seurat_obj <- compute_neighbors(
seurat_obj,
method = "hnsw",
n_neighbors = 30
)
# 5. Velocity with fewer genes
seurat_obj <- velocity(
seurat_obj,
mode = "dynamical",
n_top_genes = 1000, # Reduced from 2000
max_iter = 8, # Reduced from 10
n_cores = n_cores
)
# 6. Build velocity graph
seurat_obj <- velocity_graph(
seurat_obj,
n_neighbors = 30,
n_cores = n_cores
)
# 7. Reset backend
plan(sequential)
gc()Memory-Efficient Visualization
# Subsample for visualization
set.seed(42)
sample_idx <- sample(ncol(seurat_obj), min(5000, ncol(seurat_obj)))
# Create subsampled plot
p <- plot_velocity(seurat_obj[, sample_idx],
embedding = "umap",
n_arrows = 500)
# Save to file instead of displaying
ggsave("velocity_plot.pdf", p, width = 8, height = 6)Troubleshooting Performance Issues
Common Issues and Solutions
| Issue | Cause | Solution |
|---|---|---|
| Out of memory | Dense matrices | Use sparse matrices |
| Slow KNN | Large dataset | Use HNSW |
| Long EM runtime | Too many genes | Reduce n_top_genes |
| Worker errors | Memory per worker | Reduce workers or increase RAM |
Quick Diagnostics
# System info
Sys.info()
.Machine$sizeof.pointer # 4 = 32-bit, 8 = 64-bit
# R memory limit (Windows)
memory.limit()
# Check for data issues
sum(is.na(seurat_obj@misc$scVeloR$Ms)) # NA values
range(seurat_obj@misc$scVeloR$Ms) # Value rangesSummary
Key optimization strategies:
- Memory: Use sparse matrices, chunk processing for very large data
-
Parallelization: Use
futurepackage with appropriate workers - Algorithms: Use approximate KNN (HNSW) for large datasets
- Parameters: Reduce genes and EM iterations for speed
Session Information
sessionInfo()
#> R version 4.4.0 (2024-04-24)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] C
#>
#> time zone: Asia/Shanghai
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.1
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 jsonlite_2.0.0 dplyr_1.1.4 compiler_4.4.0
#> [5] tidyselect_1.2.1 dichromat_2.0-0.1 jquerylib_0.1.4 systemfonts_1.3.1
#> [9] scales_1.4.0 textshaping_1.0.4 yaml_2.3.12 fastmap_1.2.0
#> [13] R6_2.6.1 labeling_0.4.3 generics_0.1.4 knitr_1.51
#> [17] htmlwidgets_1.6.4 tibble_3.3.1 desc_1.4.3 bslib_0.9.0
#> [21] pillar_1.11.1 RColorBrewer_1.1-3 rlang_1.1.7 cachem_1.1.0
#> [25] xfun_0.56 fs_1.6.6 sass_0.4.10 S7_0.2.1
#> [29] otel_0.2.0 cli_3.6.5 pkgdown_2.1.3 withr_3.0.2
#> [33] magrittr_2.0.4 digest_0.6.39 grid_4.4.0 lifecycle_1.0.5
#> [37] vctrs_0.7.1 evaluate_1.0.5 glue_1.8.0 farver_2.1.2
#> [41] ragg_1.5.0 rmarkdown_2.30 tools_4.4.0 pkgconfig_2.0.3
#> [45] htmltools_0.5.9