Mathematical Framework and Algorithm
Zaoqu Liu
2026-01-26
Source:vignettes/algorithm.Rmd
algorithm.RmdIntroduction
CellProgramMapper implements a projection-based approach for mapping single-cell transcriptomic data onto reference gene expression programs (GEPs). This vignette provides a detailed explanation of the mathematical framework underlying the algorithm.
Problem Formulation
Non-Negative Matrix Factorization (NMF)
Gene expression programs are typically discovered using Non-Negative Matrix Factorization (NMF). Given an expression matrix (n cells x p genes), NMF seeks to find:
where:
- : Usage matrix (n cells x k programs)
- : Spectra matrix (k programs x p genes)
The non-negativity constraints and ensure biological interpretability.
Cell-wise Decomposition
The objective function decomposes over cells:
where is the expression vector for cell , and is its usage vector.
This allows us to solve independent Non-Negative Least Squares (NNLS) problems:
NNLS Problem
Data Preprocessing
Usage Normalization
After obtaining the raw usage matrix , rows are normalized to sum to 1:
This normalized usage represents the relative contribution of each program to a cell’s expression profile.
Computational Complexity
For a single cell with genes and programs:
- Active Set Method: per iteration, total
- Coordinate Descent: per iteration, total
where is the number of iterations.
For cells, the total complexity is or .
Example: Geometric Interpretation
The NNLS solution can be interpreted geometrically. Each cell’s expression profile is projected onto the non-negative cone spanned by the reference program spectra.
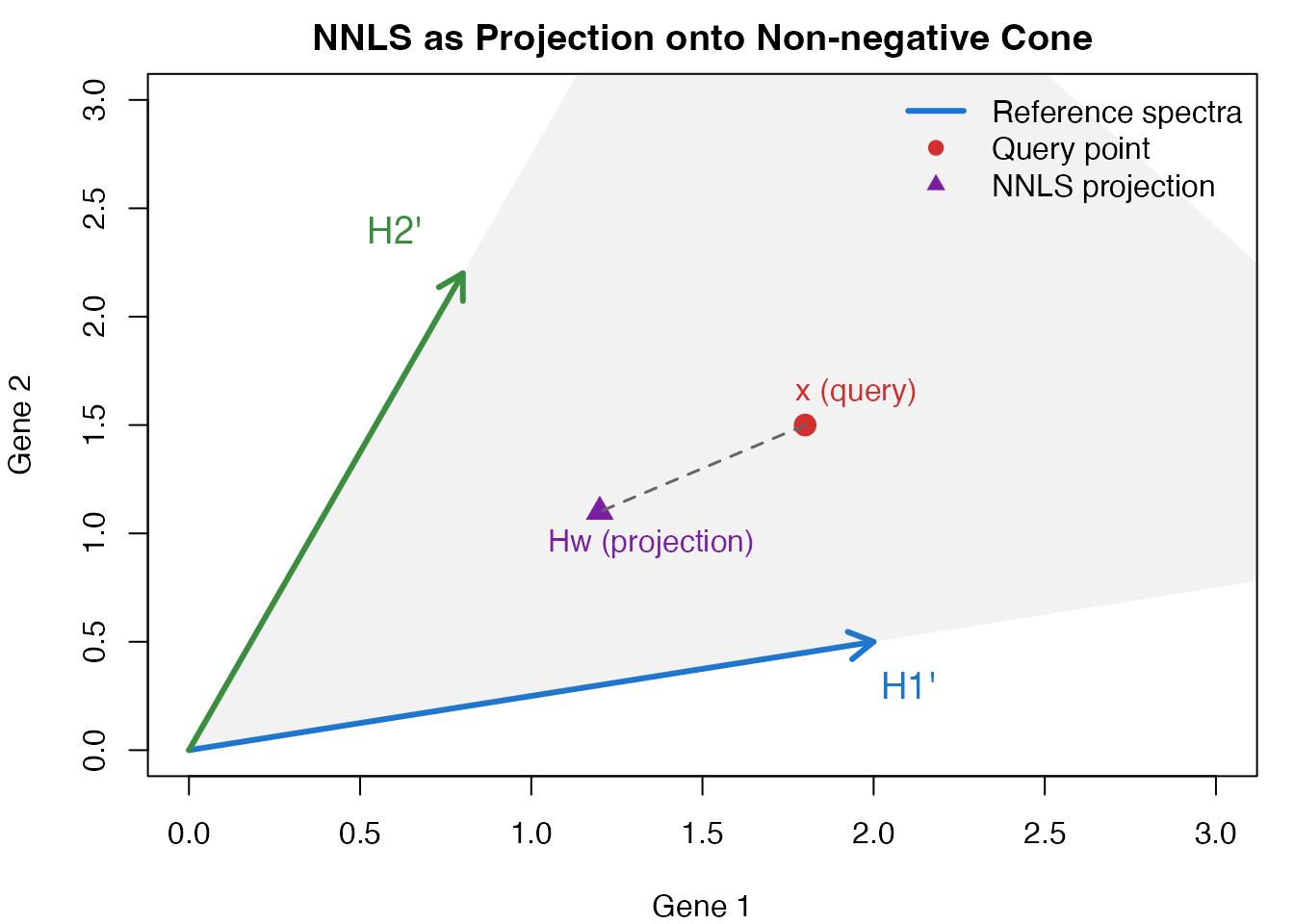
Geometric interpretation of NNLS projection
Summary
| Component | Description |
|---|---|
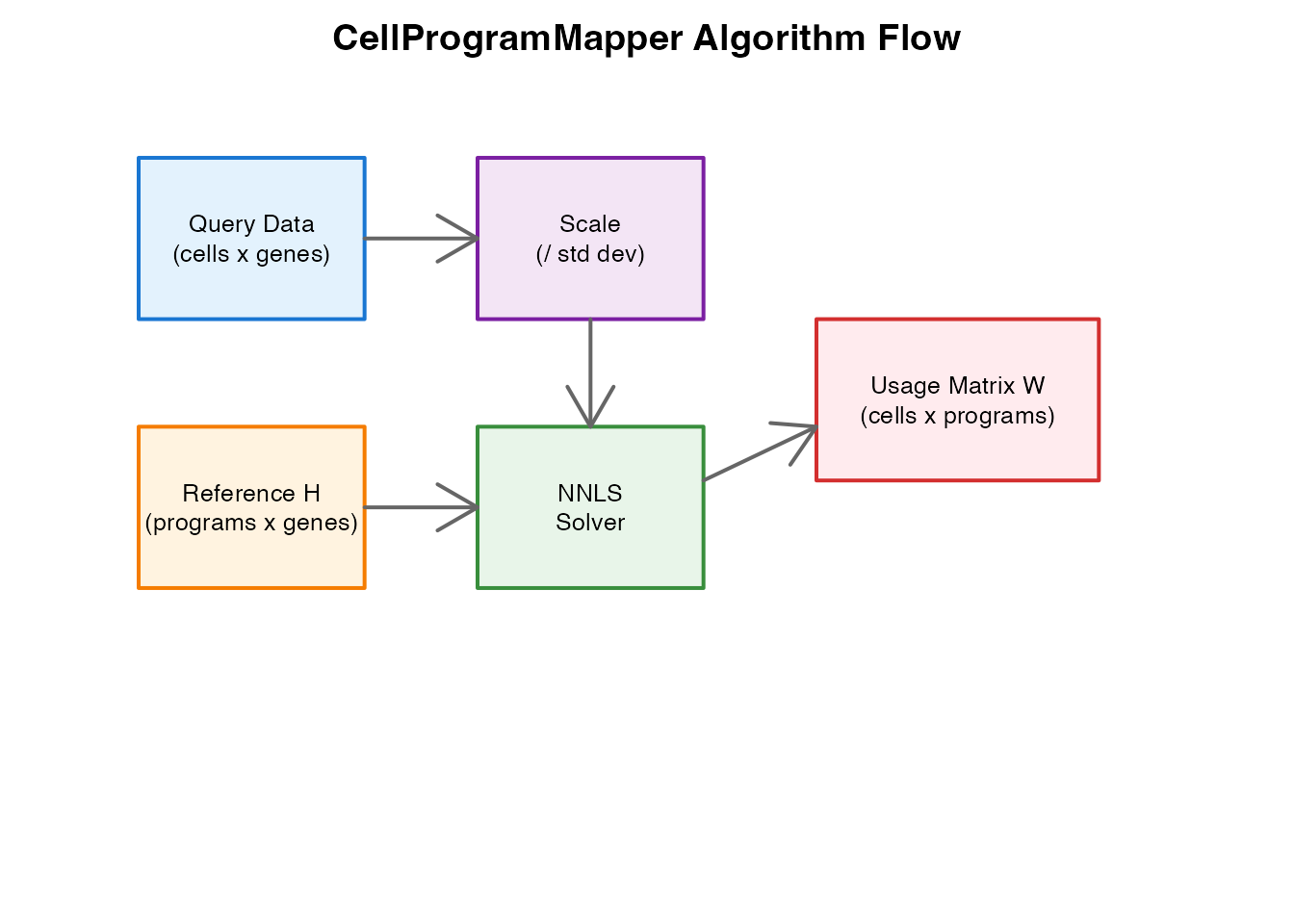
| Input | Query matrix X (cells x genes), Reference H (programs x genes) |
| Output | Usage matrix W (cells x programs) |
| Preprocessing | Scale by population standard deviation |
| Solver | NNLS (Coordinate Descent or Active Set) |
| Post-processing | Row normalization (optional) |
References
Lee DD, Seung HS (1999). Learning the parts of objects by non-negative matrix factorization. Nature 401:788-791.
Lawson CL, Hanson RJ (1974). Solving Least Squares Problems. Prentice-Hall.
Franc V, Hlavac V, Navara M (2005). Sequential Coordinate-Wise Algorithm for the Non-negative Least Squares Problem. CAIP 2005.
Session Info
sessionInfo()
#> R version 4.4.0 (2024-04-24)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] C
#>
#> time zone: Asia/Shanghai
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] digest_0.6.39 desc_1.4.3 R6_2.6.1 fastmap_1.2.0
#> [5] xfun_0.56 cachem_1.1.0 knitr_1.51 htmltools_0.5.9
#> [9] rmarkdown_2.30 lifecycle_1.0.5 cli_3.6.5 sass_0.4.10
#> [13] pkgdown_2.2.0 textshaping_1.0.4 jquerylib_0.1.4 systemfonts_1.3.1
#> [17] compiler_4.4.0 tools_4.4.0 ragg_1.5.0 bslib_0.9.0
#> [21] evaluate_1.0.5 yaml_2.3.12 otel_0.2.0 jsonlite_2.0.0
#> [25] rlang_1.1.7 fs_1.6.6 htmlwidgets_1.6.4