Introduction
This vignette demonstrates various visualization approaches for CellProgramMapper results. We cover:
- Usage matrix heatmaps
- Program activity on UMAP/t-SNE
- Score distributions
- Program correlation analysis
- Publication-ready figures
Load Example Data
library(CellProgramMapper)
# For visualization
if (!requireNamespace("ggplot2", quietly = TRUE)) {
install.packages("ggplot2")
}
if (!requireNamespace("pheatmap", quietly = TRUE)) {
install.packages("pheatmap")
}
library(ggplot2)
library(pheatmap)Simulated Example Data
For demonstration, we create simulated results:
set.seed(42)
# Simulate usage matrix (cells × programs)
n_cells <- 500
n_programs <- 8
# Create usage with structure
group <- sample(1:4, n_cells, replace = TRUE)
usage <- matrix(0, nrow = n_cells, ncol = n_programs)
# Each group has dominant programs
for (i in 1:n_cells) {
base_usage <- runif(n_programs, 0.01, 0.1)
dominant <- ((group[i] - 1) * 2 + 1):((group[i] - 1) * 2 + 2)
base_usage[dominant] <- runif(2, 0.3, 0.5)
usage[i, ] <- base_usage / sum(base_usage) # Normalize
}
colnames(usage) <- paste0("GEP", 1:n_programs)
rownames(usage) <- paste0("Cell", 1:n_cells)
# Simulate UMAP coordinates
umap <- data.frame(
UMAP1 = c(rnorm(n_cells/4, -3, 1), rnorm(n_cells/4, 3, 1),
rnorm(n_cells/4, 0, 1), rnorm(n_cells/4, 0, 1)),
UMAP2 = c(rnorm(n_cells/4, 0, 1), rnorm(n_cells/4, 0, 1),
rnorm(n_cells/4, -3, 1), rnorm(n_cells/4, 3, 1))
)
rownames(umap) <- rownames(usage)
# Simulate scores
scores <- data.frame(
Exhaustion = rowSums(usage[, c("GEP1", "GEP2")]),
Cytotoxicity = rowSums(usage[, c("GEP3", "GEP4")]),
Proliferation = rowSums(usage[, c("GEP5", "GEP6")])
)Usage Matrix Heatmap
The usage matrix shows the contribution of each GEP to each cell’s expression profile.
# Order cells by dominant program
dominant_program <- apply(usage, 1, which.max)
cell_order <- order(dominant_program, decreasing = FALSE)
# Create annotation
annotation_row <- data.frame(
DominantGEP = factor(paste0("GEP", dominant_program))
)
rownames(annotation_row) <- rownames(usage)
# Color palette
colors <- colorRampPalette(c("#f7fbff", "#08306b"))(100)
pheatmap(
usage[cell_order, ],
cluster_rows = FALSE,
cluster_cols = TRUE,
show_rownames = FALSE,
annotation_row = annotation_row[cell_order, , drop = FALSE],
color = colors,
main = "Cell × GEP Usage Matrix",
fontsize = 10
)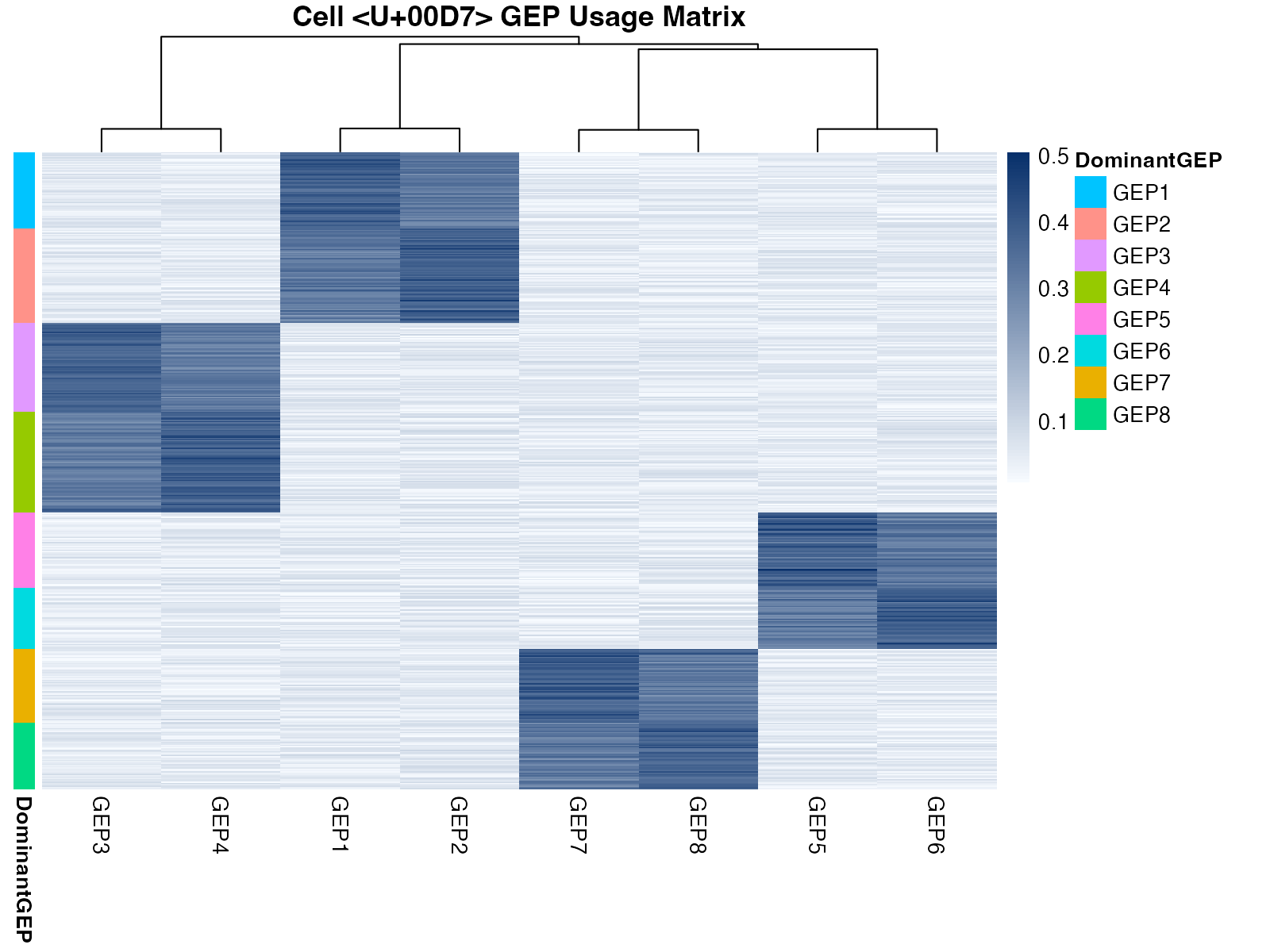
Usage matrix heatmap showing GEP contributions across cells
Program Activity on UMAP
Visualize GEP activity overlaid on dimensionality reduction.
# Combine data
plot_data <- cbind(umap, usage)
# Create multi-panel plot
par(mfrow = c(2, 4), mar = c(4, 4, 2, 2))
for (gep in colnames(usage)) {
colors <- colorRampPalette(c("#f7f7f7", "#d73027"))(100)
vals <- plot_data[[gep]]
col_idx <- cut(vals, breaks = 100, labels = FALSE)
plot(plot_data$UMAP1, plot_data$UMAP2,
col = colors[col_idx], pch = 16, cex = 0.6,
xlab = "UMAP1", ylab = "UMAP2",
main = gep)
}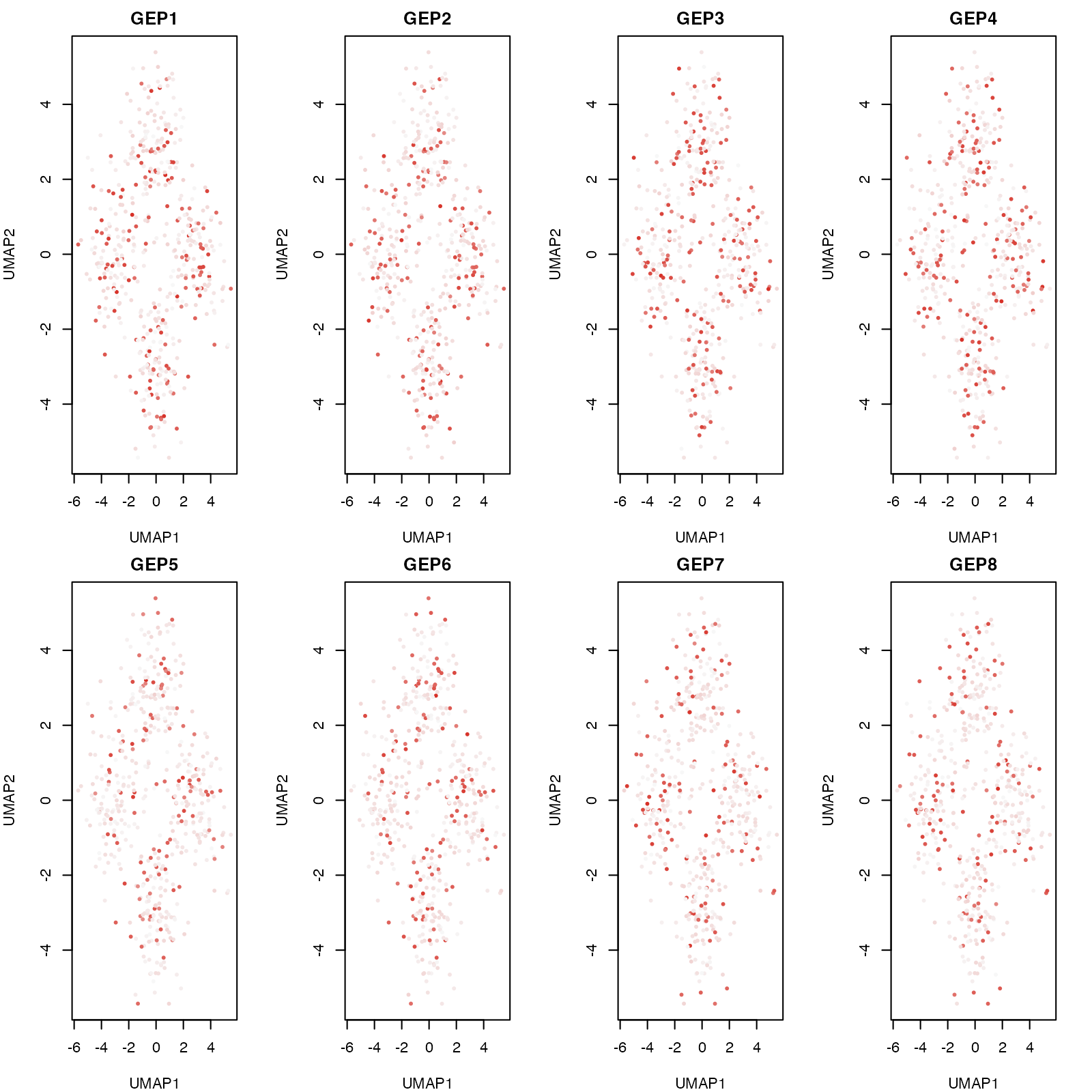
GEP activity visualized on UMAP coordinates
Score Distributions
Violin Plots
# Create data for violin plot
scores$Group <- factor(paste0("Group", group))
# Reshape for ggplot
library(reshape2)
#> Warning: package 'reshape2' was built under R version 4.4.1
scores_long <- melt(scores, id.vars = "Group",
variable.name = "Score", value.name = "Value")
ggplot(scores_long, aes(x = Group, y = Value, fill = Score)) +
geom_violin(alpha = 0.7, scale = "width") +
geom_boxplot(width = 0.1, fill = "white", alpha = 0.8) +
facet_wrap(~Score, scales = "free_y") +
theme_minimal() +
labs(title = "Score Distributions by Group",
x = "Cell Group", y = "Score Value") +
theme(legend.position = "none",
strip.text = element_text(face = "bold", size = 12))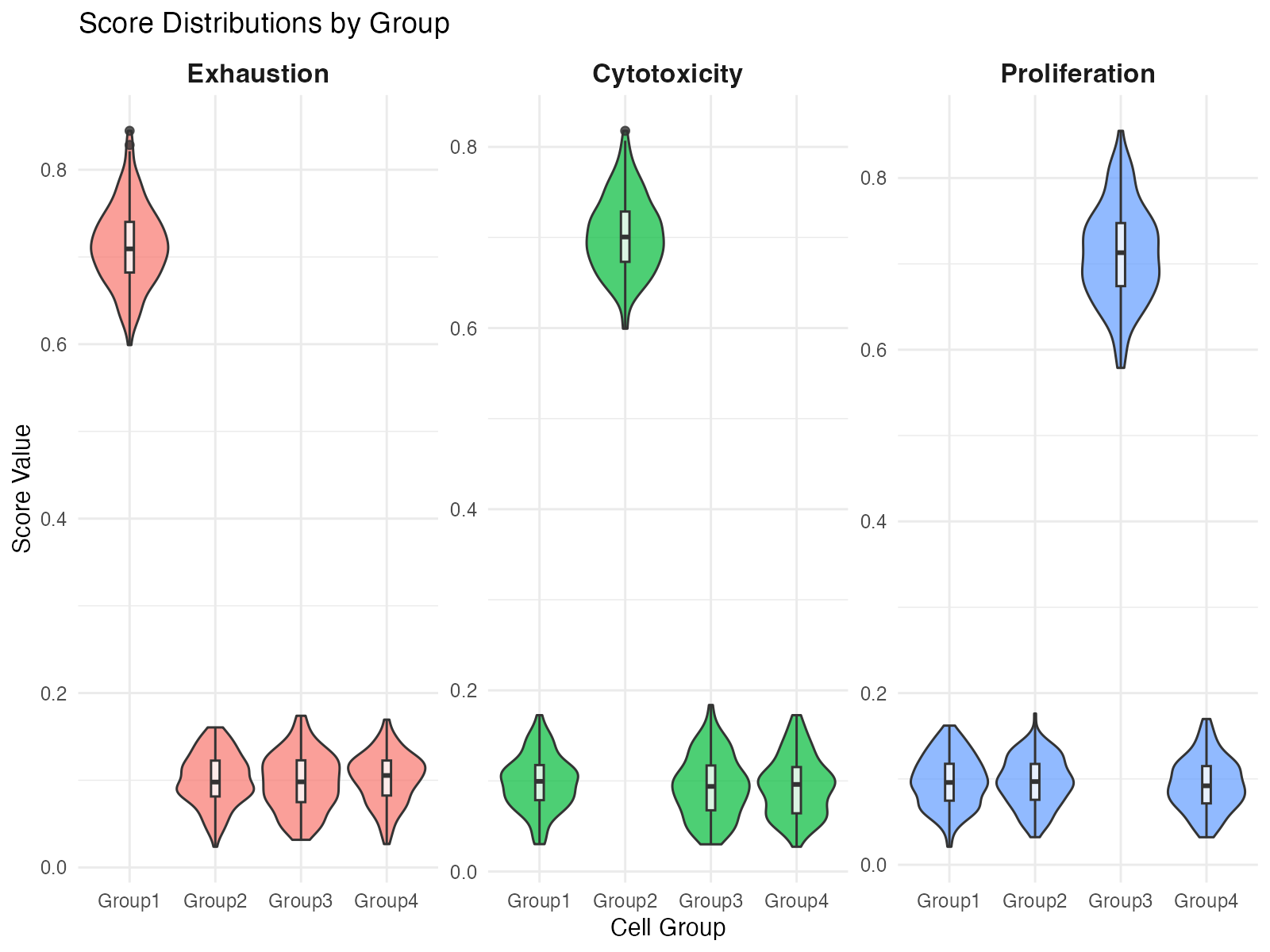
Score distributions across groups
Ridge Plot
# Simple ridge plot using base R
par(mfrow = c(3, 1), mar = c(3, 4, 2, 1))
for (score_name in c("Exhaustion", "Cytotoxicity", "Proliferation")) {
# Get density for each group
densities <- lapply(1:4, function(g) {
d <- density(scores[[score_name]][scores$Group == paste0("Group", g)])
data.frame(x = d$x, y = d$y, group = g)
})
# Plot
x_range <- range(sapply(densities, function(d) range(d$x)))
y_max <- max(sapply(densities, function(d) max(d$y))) * 1.2
plot(NULL, xlim = x_range, ylim = c(-0.5, y_max * 4.5),
xlab = score_name, ylab = "", yaxt = "n",
main = paste0(score_name, " Score"))
cols <- c("#e41a1c", "#377eb8", "#4daf4a", "#984ea3")
for (i in 1:4) {
d <- densities[[i]]
offset <- (i - 1) * y_max
polygon(c(d$x, rev(d$x)),
c(d$y + offset, rep(offset, length(d$x))),
col = adjustcolor(cols[i], alpha.f = 0.6),
border = cols[i])
}
axis(2, at = (0:3) * y_max + y_max/2,
labels = paste0("Group", 1:4), las = 1)
}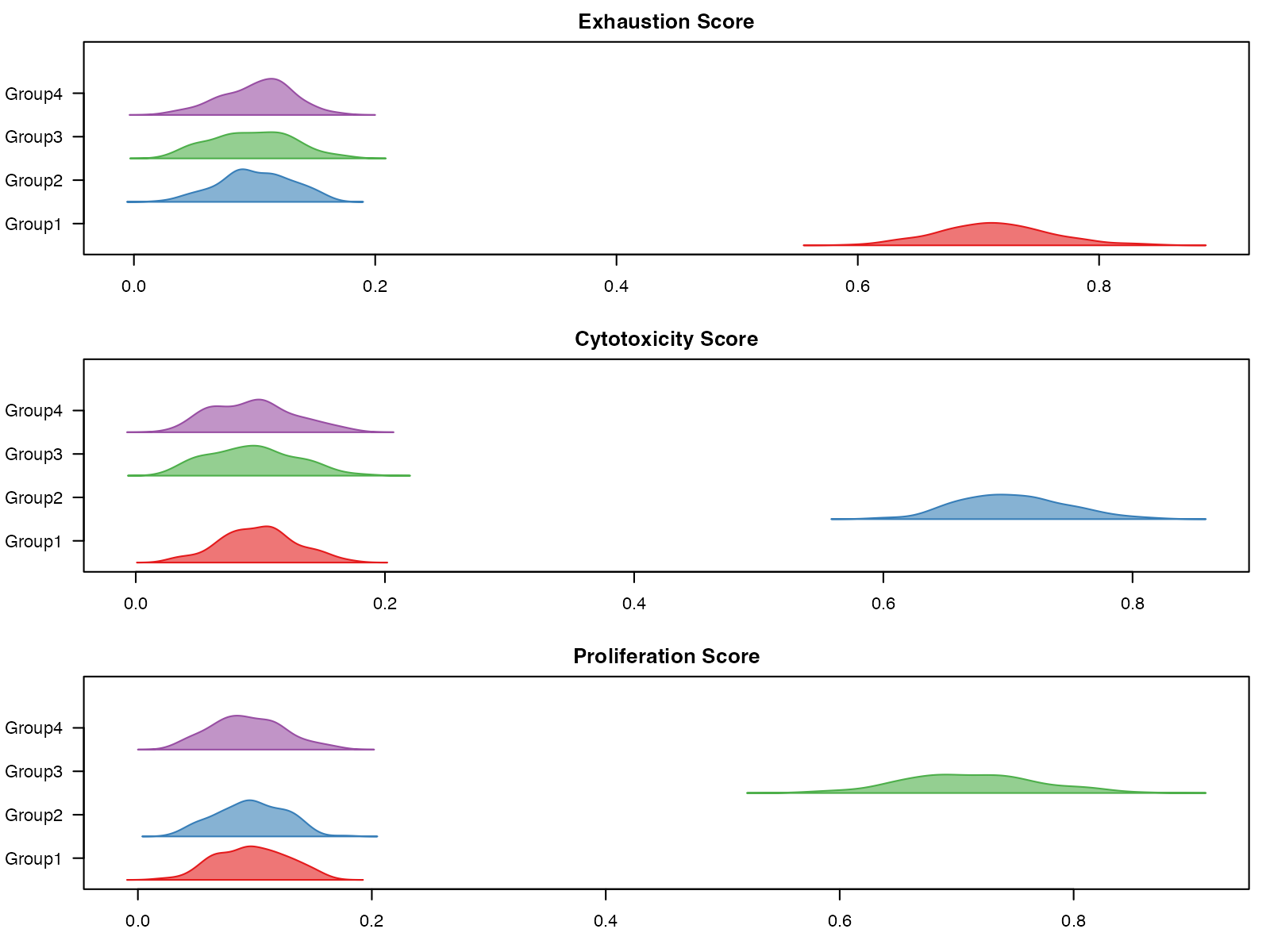
Ridge plot of score distributions
Program Correlation Analysis
Correlation Heatmap
# Compute correlation
usage_cor <- cor(usage)
# Custom color palette
cor_colors <- colorRampPalette(c("#2166ac", "#f7f7f7", "#b2182b"))(100)
pheatmap(
usage_cor,
color = cor_colors,
breaks = seq(-1, 1, length.out = 101),
display_numbers = TRUE,
number_format = "%.2f",
fontsize_number = 8,
main = "GEP Correlation Matrix"
)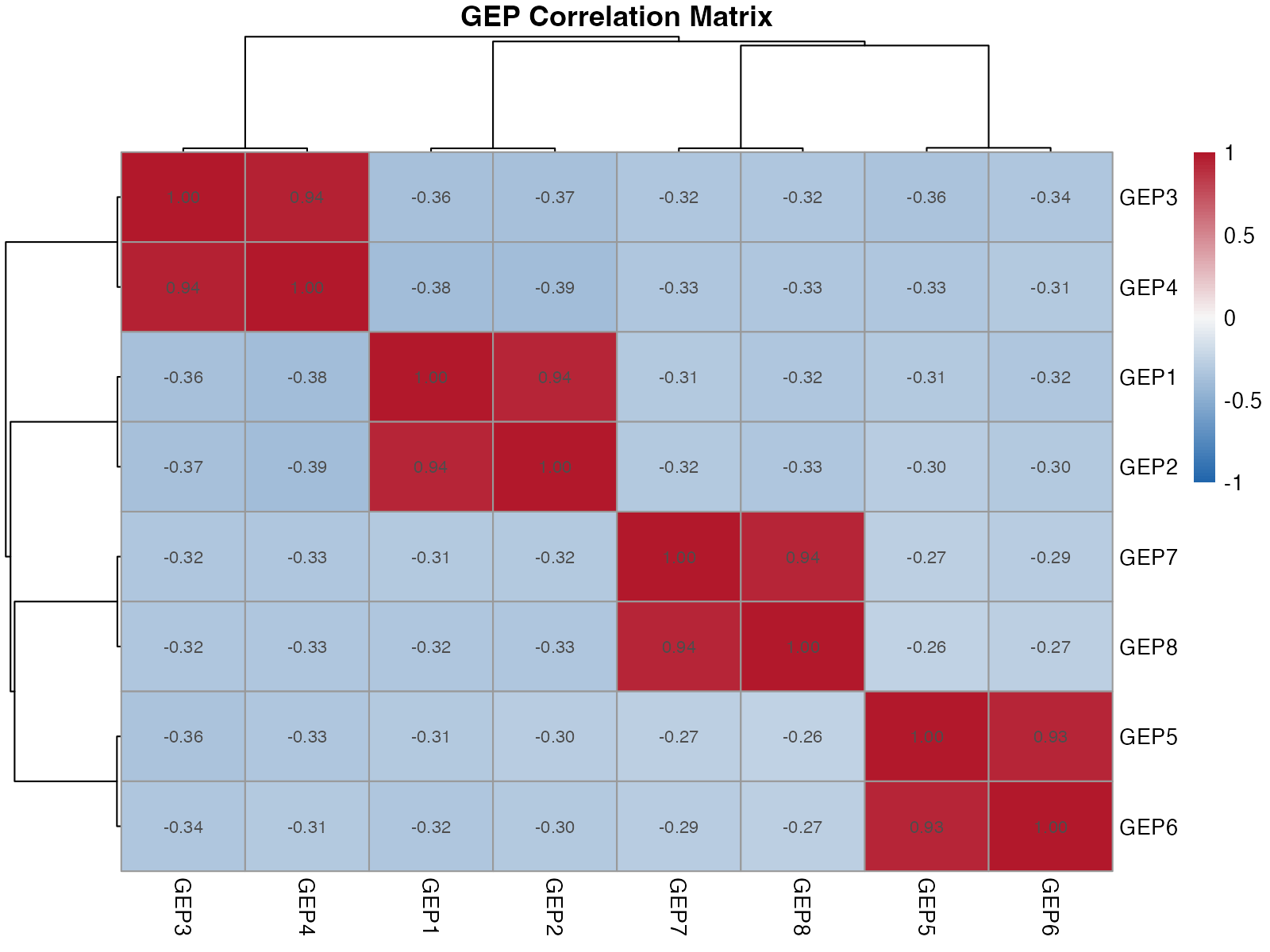
Correlation between GEPs
Network Visualization
# Simple network plot
par(mar = c(1, 1, 2, 1))
plot(NULL, xlim = c(-1.5, 1.5), ylim = c(-1.5, 1.5),
xlab = "", ylab = "", axes = FALSE,
main = "GEP Correlation Network")
# Arrange nodes in circle
n <- ncol(usage)
angles <- seq(0, 2*pi, length.out = n + 1)[1:n]
x_pos <- cos(angles)
y_pos <- sin(angles)
# Draw edges (correlations > 0.3)
for (i in 1:(n-1)) {
for (j in (i+1):n) {
r <- usage_cor[i, j]
if (abs(r) > 0.3) {
col <- if (r > 0) "#b2182b" else "#2166ac"
lwd <- abs(r) * 3
segments(x_pos[i], y_pos[i], x_pos[j], y_pos[j],
col = adjustcolor(col, alpha.f = 0.5), lwd = lwd)
}
}
}
# Draw nodes
points(x_pos, y_pos, pch = 19, cex = 4, col = "#4a4a4a")
text(x_pos, y_pos, colnames(usage), col = "white", cex = 0.7, font = 2)
legend("topright",
legend = c("Positive corr.", "Negative corr."),
col = c("#b2182b", "#2166ac"),
lwd = 2, bty = "n")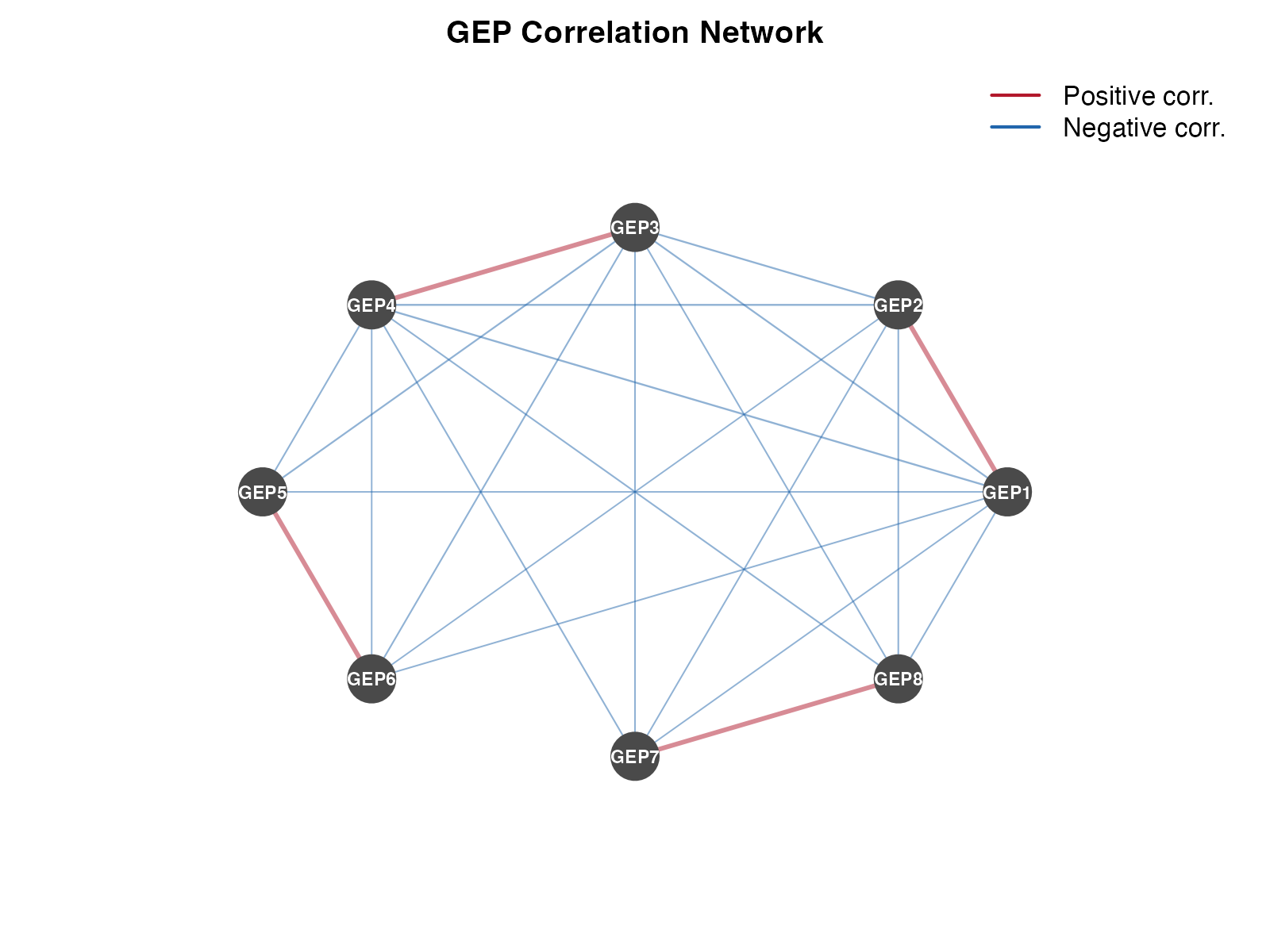
GEP correlation network
Publication-Ready Figures
Combined Panel Figure
par(mfrow = c(2, 2), mar = c(4, 4, 3, 2))
# Panel A: UMAP colored by dominant GEP
dominant <- factor(paste0("GEP", apply(usage, 1, which.max)))
cols <- rainbow(8, alpha = 0.7)
plot(umap$UMAP1, umap$UMAP2, col = cols[dominant], pch = 16,
xlab = "UMAP1", ylab = "UMAP2", main = "A. Dominant GEP")
legend("topright", levels(dominant), col = cols, pch = 16, cex = 0.6, ncol = 2)
# Panel B: Usage bar plot (top cells)
top_cells <- head(order(usage[, "GEP1"], decreasing = TRUE), 20)
barplot(t(usage[top_cells, ]),
col = cols, border = NA, las = 2,
xlab = "", ylab = "Usage",
main = "B. Top GEP1 Cells")
# Panel C: Score comparison
boxplot(Exhaustion ~ Group, data = scores,
col = c("#e41a1c", "#377eb8", "#4daf4a", "#984ea3"),
xlab = "Group", ylab = "Exhaustion Score",
main = "C. Exhaustion by Group")
# Panel D: Program correlation
image(1:8, 1:8, usage_cor[8:1, ],
col = cor_colors,
xlab = "", ylab = "", axes = FALSE,
main = "D. GEP Correlations")
axis(1, at = 1:8, labels = colnames(usage), las = 2, cex.axis = 0.8)
axis(2, at = 1:8, labels = rev(colnames(usage)), las = 1, cex.axis = 0.8)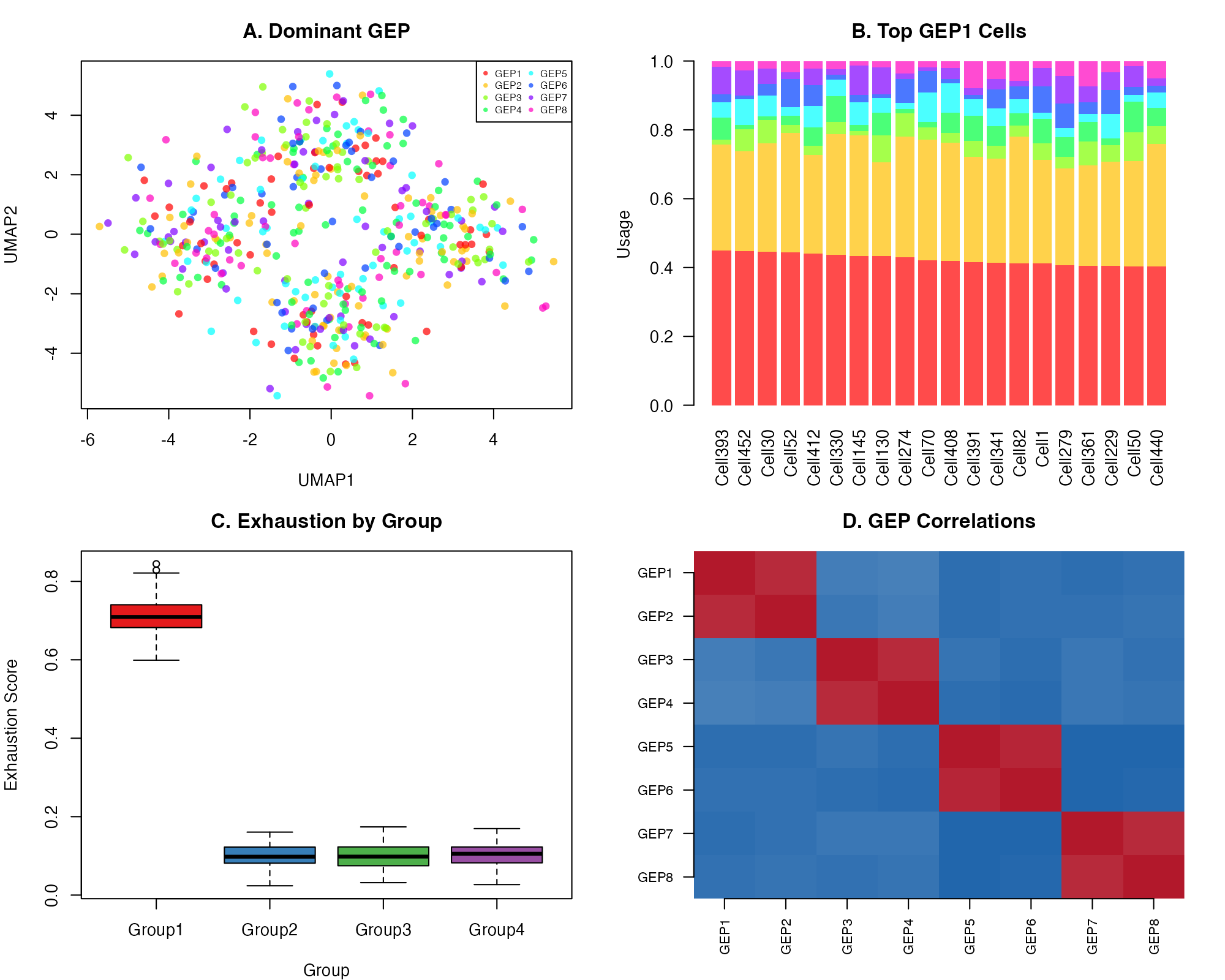
Publication-ready combined visualization
Saving High-Resolution Figures
# For publication (300 DPI)
pdf("figures/usage_heatmap.pdf", width = 8, height = 6)
pheatmap(usage[cell_order, ], ...)
dev.off()
# For presentations (PNG)
png("figures/umap_panel.png", width = 2400, height = 1800, res = 300)
# ... plotting code ...
dev.off()
# For vector graphics (SVG)
svg("figures/correlation_network.svg", width = 8, height = 6)
# ... plotting code ...
dev.off()Integration with Seurat
When working with Seurat objects, use built-in visualization functions:
# Add results to Seurat
seurat_obj <- add_results_to_seurat(seurat_obj, result)
# Seurat's DimPlot
Seurat::DimPlot(seurat_obj, group.by = "Multinomial_Label",
label = TRUE, repel = TRUE)
# Seurat's FeaturePlot for GEP usage
Seurat::FeaturePlot(seurat_obj,
features = c("GEP1", "GEP2", "GEP3", "GEP4"),
ncol = 2)
# Seurat's VlnPlot for scores
Seurat::VlnPlot(seurat_obj,
features = "Exhaustion_Score",
group.by = "seurat_clusters")Tips for Effective Visualization
- Color choice: Use colorblind-friendly palettes
- Scale bars: Always include when showing continuous values
- Labels: Make text readable (≥8pt for publication)
- Consistency: Use same color scheme across related figures
- Simplicity: Avoid cluttering with unnecessary elements
Session Info
sessionInfo()
#> R version 4.4.0 (2024-04-24)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] C
#>
#> time zone: Asia/Shanghai
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] reshape2_1.4.5 pheatmap_1.0.13 ggplot2_4.0.1
#> [4] CellProgramMapper_1.0.0
#>
#> loaded via a namespace (and not attached):
#> [1] rappdirs_0.3.4 sass_0.4.10 future_1.69.0
#> [4] generics_0.1.4 stringi_1.8.7 lattice_0.22-7
#> [7] listenv_0.10.0 digest_0.6.39 magrittr_2.0.4
#> [10] evaluate_1.0.5 grid_4.4.0 RColorBrewer_1.1-3
#> [13] fastmap_1.2.0 plyr_1.8.9 jsonlite_2.0.0
#> [16] Matrix_1.7-4 scales_1.4.0 codetools_0.2-20
#> [19] textshaping_1.0.4 jquerylib_0.1.4 cli_3.6.5
#> [22] rlang_1.1.7 parallelly_1.46.1 future.apply_1.20.1
#> [25] withr_3.0.2 cachem_1.1.0 yaml_2.3.12
#> [28] otel_0.2.0 tools_4.4.0 parallel_4.4.0
#> [31] dplyr_1.1.4 globals_0.18.0 curl_7.0.0
#> [34] vctrs_0.7.1 R6_2.6.1 lifecycle_1.0.5
#> [37] stringr_1.6.0 fs_1.6.6 htmlwidgets_1.6.4
#> [40] ragg_1.5.0 pkgconfig_2.0.3 desc_1.4.3
#> [43] pkgdown_2.2.0 bslib_0.9.0 pillar_1.11.1
#> [46] gtable_0.3.6 data.table_1.18.0 glue_1.8.0
#> [49] Rcpp_1.1.1 systemfonts_1.3.1 tidyselect_1.2.1
#> [52] xfun_0.56 tibble_3.3.1 knitr_1.51
#> [55] dichromat_2.0-0.1 farver_2.1.2 htmltools_0.5.9
#> [58] labeling_0.4.3 rmarkdown_2.30 compiler_4.4.0
#> [61] S7_0.2.1