NNLS Solver Implementation Details
Zaoqu Liu
2026-01-26
Source:vignettes/nnls-solver.Rmd
nnls-solver.RmdOverview
CellProgramMapper implements two algorithms for solving the Non-Negative Least Squares (NNLS) problem:
- Coordinate Descent (default, faster)
- Active Set Method (Lawson-Hanson algorithm)
Both solvers are implemented in C++ using RcppArmadillo for optimal performance.
Active Set Method (Lawson-Hanson)
Algorithm Description
The active set method partitions variables into two sets:
- P (Positive set): Variables that can be positive
- Z (Zero set): Variables constrained to zero
Algorithm: Lawson-Hanson NNLS
Input: A (m x n matrix), b (m-vector)
Output: x (n-vector) with x >= 0
1. Initialize: x = 0, P = empty, Z = {1,...,n}
2. Compute gradient: w = A'(b - Ax)
3. While Z is not empty and max(w[Z]) > 0:
a. Move index of max(w[Z]) from Z to P
b. Inner loop:
- Solve unconstrained LS on P: z[P] = argmin ||A[:,P]z - b||^2
- While any z[P] <= 0:
* Compute step alpha to boundary
* Update x = x + alpha(z - x)
* Move zero indices from P to Z
c. Update gradient: w = A'(b - Ax)
4. Return xConvergence Properties
- Finite termination: Guaranteed to terminate in at most iterations
- Exact solution: Returns the globally optimal solution
- Numerical stability: Uses QR decomposition for least squares subproblems
Implementation
// Simplified C++ implementation (actual code in src/nnls_solver.cpp)
arma::vec nnls_single(const arma::mat& A, const arma::vec& b,
int max_iter, double tol) {
int n = A.n_cols;
arma::vec x = arma::zeros(n);
arma::vec w = A.t() * b; // Initial gradient
arma::uvec P_set, Z_set = arma::regspace<arma::uvec>(0, n-1);
while (Z_set.n_elem > 0 && iter < max_iter) {
// Find index with max positive gradient in Z
arma::uword t = Z_set(w.elem(Z_set).index_max());
if (w(t) <= tol) break;
// Move t from Z to P
P_set = join_cols(P_set, {t});
Z_set.shed_row(find_idx);
// Inner loop: solve unconstrained subproblem
while (true) {
arma::vec z_P = solve(A.cols(P_set), b);
if (all(z_P > 0)) {
x.elem(P_set) = z_P;
break;
}
// Handle negative values...
}
w = A.t() * (b - A * x);
}
return x;
}Coordinate Descent Method
Algorithm Description
Coordinate descent optimizes one variable at a time while keeping others fixed.
For NNLS, we work with the normal equations:
where and .
Algorithm: Coordinate Descent NNLS
Input: Q = A'A, c = A'b
Output: x (n-vector) with x >= 0
1. Initialize: x = 0, g = -c (gradient at x=0)
2. Repeat until convergence:
For j = 1 to n:
a. Compute update: delta = -g[j] / Q[j,j]
b. Project: x_new = max(0, x[j] + delta)
c. Update gradient: g = g + (x_new - x[j]) * Q[:,j]
d. x[j] = x_new
3. Return xAdvantages
- Simplicity: Easy to implement and understand
- Efficiency: Each iteration is for gradient update
- Precomputation: computed once for batch processing
Implementation
// Simplified C++ implementation
arma::vec nnls_coord_descent(const arma::mat& AtA, const arma::vec& Atb,
int max_iter, double tol) {
int n = AtA.n_cols;
arma::vec x = arma::zeros(n);
arma::vec grad = -Atb; // Gradient at x=0
for (int iter = 0; iter < max_iter; iter++) {
double max_change = 0;
for (int j = 0; j < n; j++) {
if (AtA(j,j) < 1e-12) continue; // Skip zero diagonal
double old_xj = x(j);
double new_xj = std::max(0.0, old_xj - grad(j) / AtA(j,j));
if (new_xj != old_xj) {
double delta = new_xj - old_xj;
x(j) = new_xj;
grad += delta * AtA.col(j); // Efficient gradient update
max_change = std::max(max_change, std::abs(delta));
}
}
if (max_change < tol) break;
}
return x;
}Comparison
| Aspect | Active Set | Coordinate Descent |
|---|---|---|
| Time per iteration | O(k^3) | O(k^2) |
| Iterations to converge | Usually fewer | Usually more |
| Memory | O(p*k) | O(k^2) |
| Convergence guarantee | Finite | Linear |
| Best for | Small k | Large k |
Benchmark
library(CellProgramMapper)
#> CellProgramMapper v1.0.0
#> Map single cells to reference gene expression programs
#> GitHub: https://github.com/Zaoqu-Liu/CellProgramMapper
# Benchmark function using CellProgramMapper
benchmark_solver <- function(n_cells, n_genes, n_programs, method) {
set.seed(42)
X <- matrix(abs(rnorm(n_cells * n_genes)), nrow = n_cells)
colnames(X) <- paste0("Gene", 1:n_genes)
H <- matrix(abs(rnorm(n_programs * n_genes)), nrow = n_programs)
colnames(H) <- paste0("Gene", 1:n_genes)
rownames(H) <- paste0("GEP", 1:n_programs)
start_time <- Sys.time()
result <- CellProgramMapper(
query = X,
reference = H,
method = method,
verbose = FALSE
)
end_time <- Sys.time()
as.numeric(difftime(end_time, start_time, units = "secs"))
}
# Run benchmarks
n_cells_vec <- c(100, 500, 1000)
n_genes <- 200
n_programs <- 10
results <- data.frame()
for (n_cells in n_cells_vec) {
for (method in c("cd", "active_set")) {
time_taken <- benchmark_solver(n_cells, n_genes, n_programs, method)
results <- rbind(results, data.frame(
n_cells = n_cells,
method = method,
time = time_taken
))
}
}
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
#> Warning: Query data does not appear to be integer counts. For best results,
#> provide raw UMI/read counts.
# Plot
par(mar = c(4, 4, 2, 1))
cd_times <- results$time[results$method == "cd"]
as_times <- results$time[results$method == "active_set"]
barplot(rbind(cd_times, as_times), beside = TRUE,
names.arg = n_cells_vec,
col = c("#1976d2", "#388e3c"),
xlab = "Number of cells",
ylab = "Time (seconds)",
main = "NNLS Solver Performance")
legend("topleft", legend = c("Coordinate Descent", "Active Set"),
fill = c("#1976d2", "#388e3c"), bty = "n")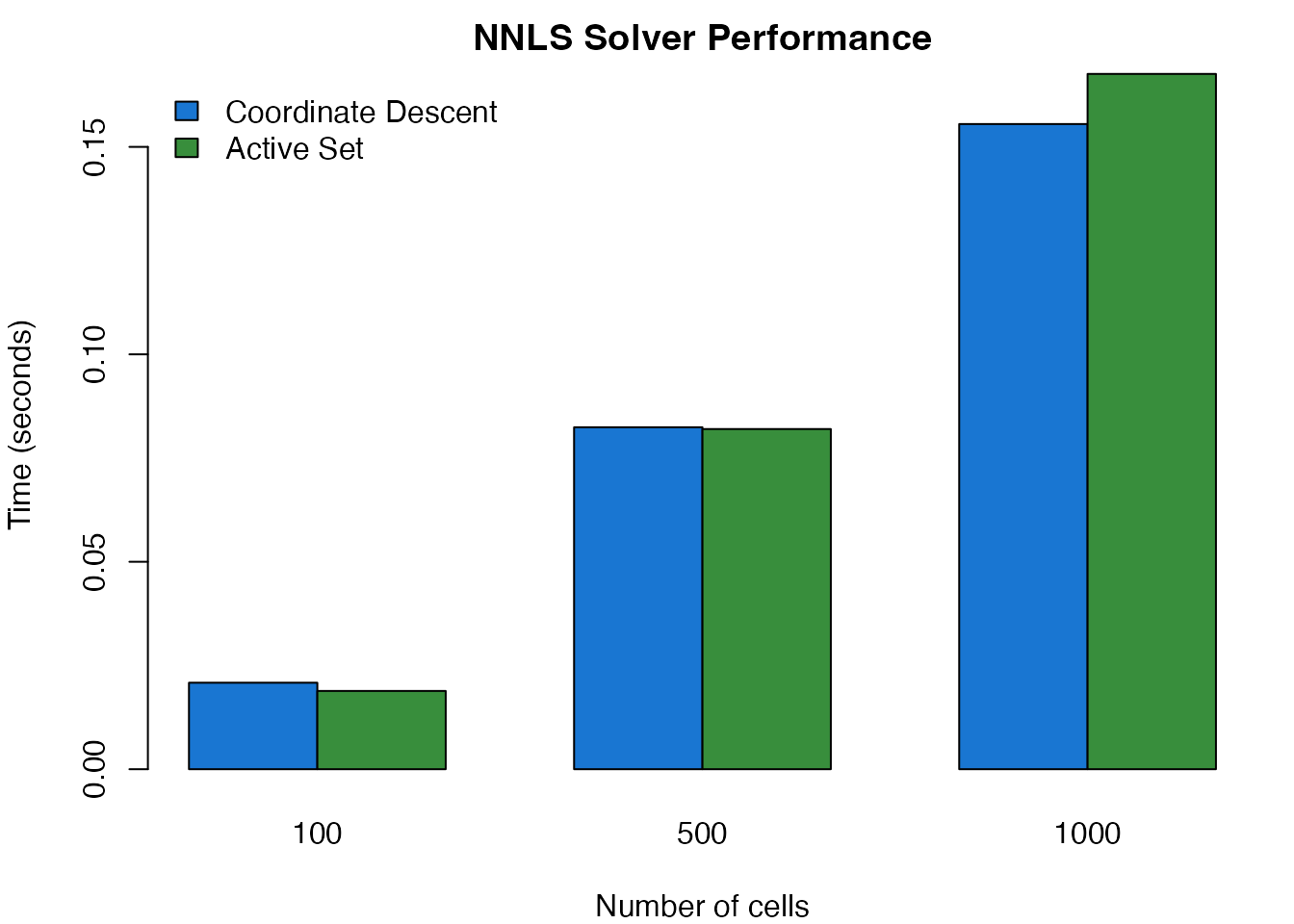
Solver performance comparison
Choosing a Solver
Use Coordinate Descent (default) when:
- Processing large numbers of cells (>1000)
- Number of programs is moderate (5-50)
- Speed is the priority
Use Active Set when:
- High numerical precision is required
- Number of programs is small (<10)
- Guaranteed finite convergence is important
# Use coordinate descent (default)
result <- CellProgramMapper(query, reference, method = "cd")
# Use active set method
result <- CellProgramMapper(query, reference, method = "active_set")Numerical Considerations
Tolerance Parameter
The tol parameter controls convergence:
- Smaller values: more iterations, higher precision
- Larger values: fewer iterations, faster but less precise
- Default:
1e-8(good balance)
Session Info
sessionInfo()
#> R version 4.4.0 (2024-04-24)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] C
#>
#> time zone: Asia/Shanghai
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] CellProgramMapper_1.0.0
#>
#> loaded via a namespace (and not attached):
#> [1] cli_3.6.5 knitr_1.51 rlang_1.1.7
#> [4] xfun_0.56 otel_0.2.0 textshaping_1.0.4
#> [7] data.table_1.18.0 jsonlite_2.0.0 future.apply_1.20.1
#> [10] listenv_0.10.0 htmltools_0.5.9 ragg_1.5.0
#> [13] sass_0.4.10 rappdirs_0.3.4 rmarkdown_2.30
#> [16] grid_4.4.0 evaluate_1.0.5 jquerylib_0.1.4
#> [19] fastmap_1.2.0 yaml_2.3.12 lifecycle_1.0.5
#> [22] compiler_4.4.0 codetools_0.2-20 fs_1.6.6
#> [25] Rcpp_1.1.1 htmlwidgets_1.6.4 future_1.69.0
#> [28] systemfonts_1.3.1 lattice_0.22-7 digest_0.6.39
#> [31] R6_2.6.1 parallelly_1.46.1 parallel_4.4.0
#> [34] curl_7.0.0 bslib_0.9.0 Matrix_1.7-4
#> [37] tools_4.4.0 globals_0.18.0 pkgdown_2.2.0
#> [40] cachem_1.1.0 desc_1.4.3