Overview
This tutorial covers the communication analysis step, where production and sensing scores are combined to quantify cell-cell communication via metabolites.
library(scMetaLink)
library(Matrix)
# Load and prepare data
data(crc_example)
obj <- createScMetaLink(crc_expr, crc_meta, "cell_type")
obj <- inferProduction(obj, verbose = FALSE)
obj <- inferSensing(obj, verbose = FALSE)Understanding Communication Scores
Communication from sender to receiver via metabolite:
Sender Cell ──[Production]──▶ Metabolite ──[Sensing]──▶ Receiver Cell
│ │ │
│ │ │
▼ ▼ ▼
MPP(m,s) -----> Comm(s->r,m) <----- MSC(m,r)The computeCommunication() Function
obj <- computeCommunication(
obj,
method = "geometric", # How to combine production and sensing
min_production = 0.1, # Minimum production threshold
min_sensing = 0.1, # Minimum sensing threshold
population.size = FALSE, # Weight by cell type abundance
n_permutations = 100, # Permutations for significance
n_cores = 1, # Parallel cores
seed = 42, # Reproducibility
verbose = TRUE
)
#> | | | 0% | |= | 1% | |= | 2% | |== | 3% | |=== | 4% | |==== | 5% | |==== | 6% | |===== | 7% | |====== | 8% | |====== | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 13% | |========== | 14% | |========== | 15% | |=========== | 16% | |============ | 17% | |============= | 18% | |============= | 19% | |============== | 20% | |=============== | 21% | |=============== | 22% | |================ | 23% | |================= | 24% | |================== | 25% | |================== | 26% | |=================== | 27% | |==================== | 28% | |==================== | 29% | |===================== | 30% | |====================== | 31% | |====================== | 32% | |======================= | 33% | |======================== | 34% | |======================== | 35% | |========================= | 36% | |========================== | 37% | |=========================== | 38% | |=========================== | 39% | |============================ | 40% | |============================= | 41% | |============================= | 42% | |============================== | 43% | |=============================== | 44% | |================================ | 45% | |================================ | 46% | |================================= | 47% | |================================== | 48% | |================================== | 49% | |=================================== | 50% | |==================================== | 51% | |==================================== | 52% | |===================================== | 53% | |====================================== | 54% | |====================================== | 55% | |======================================= | 56% | |======================================== | 57% | |========================================= | 58% | |========================================= | 59% | |========================================== | 60% | |=========================================== | 61% | |=========================================== | 62% | |============================================ | 63% | |============================================= | 64% | |============================================== | 65% | |============================================== | 66% | |=============================================== | 67% | |================================================ | 68% | |================================================ | 69% | |================================================= | 70% | |================================================== | 71% | |================================================== | 72% | |=================================================== | 73% | |==================================================== | 74% | |==================================================== | 75% | |===================================================== | 76% | |====================================================== | 77% | |======================================================= | 78% | |======================================================= | 79% | |======================================================== | 80% | |========================================================= | 81% | |========================================================= | 82% | |========================================================== | 83% | |=========================================================== | 84% | |============================================================ | 85% | |============================================================ | 86% | |============================================================= | 87% | |============================================================== | 88% | |============================================================== | 89% | |=============================================================== | 90% | |================================================================ | 91% | |================================================================ | 92% | |================================================================= | 93% | |================================================================== | 94% | |================================================================== | 95% | |=================================================================== | 96% | |==================================================================== | 97% | |===================================================================== | 98% | |===================================================================== | 99% | |======================================================================| 100%Parameter Deep Dive
1. method: Combining Production and Sensing
Three methods to compute communication score:
# Run with different methods
obj_geom <- computeCommunication(obj, method = "geometric", n_permutations = 0, verbose = FALSE)
obj_prod <- computeCommunication(obj, method = "product", n_permutations = 0, verbose = FALSE)
obj_harm <- computeCommunication(obj, method = "harmonic", n_permutations = 0, verbose = FALSE)
# Compare total communication strength
par(mfrow = c(1, 3))
total_geom <- apply(obj_geom@communication_scores, c(1, 2), sum)
total_prod <- apply(obj_prod@communication_scores, c(1, 2), sum)
total_harm <- apply(obj_harm@communication_scores, c(1, 2), sum)
image(total_geom,
main = "Geometric: sqrt(PxS)", col = hcl.colors(50, "YlOrRd"),
axes = FALSE
)
image(total_prod,
main = "Product: PxS", col = hcl.colors(50, "YlOrRd"),
axes = FALSE
)
image(total_harm,
main = "Harmonic: 2PS/(P+S)", col = hcl.colors(50, "YlOrRd"),
axes = FALSE
)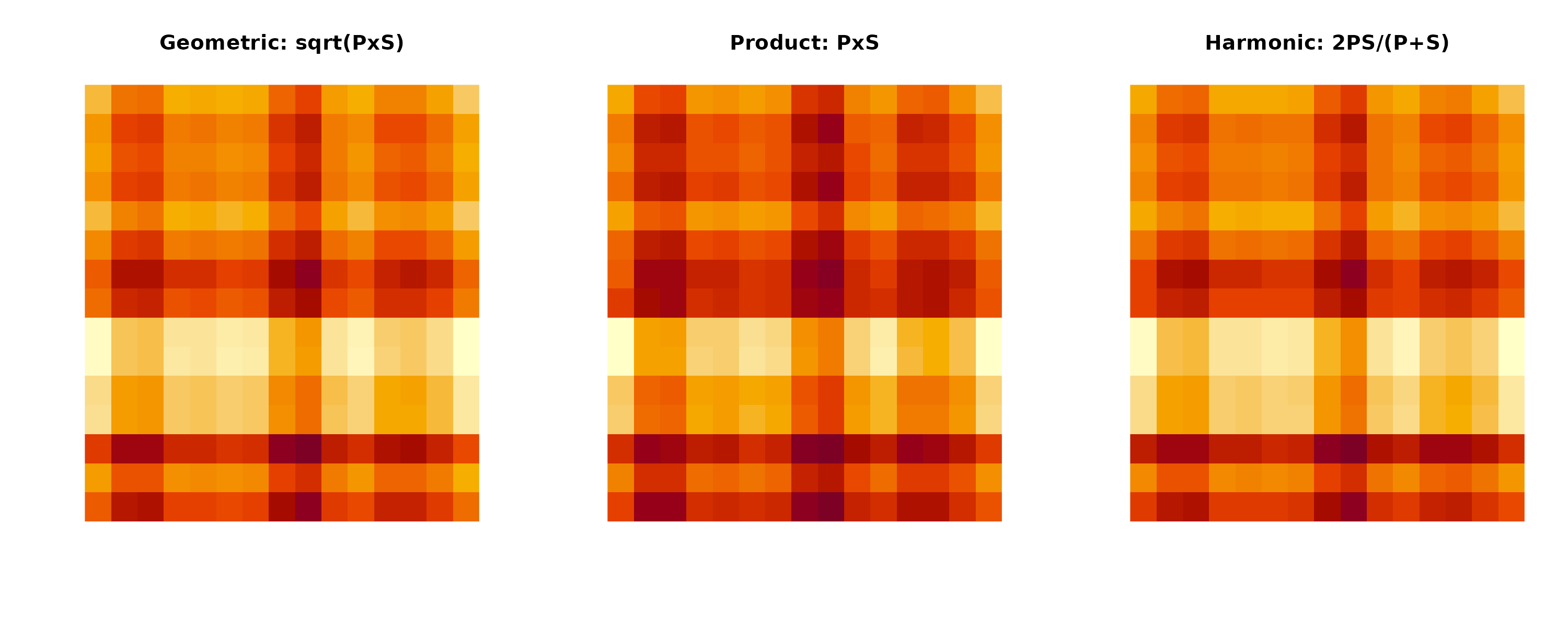
Figure 1: Communication Score Method Comparison. Heatmaps showing total communication strength between cell types using different aggregation methods. Geometric mean (default) provides balanced weighting, product emphasizes strong bilateral signals, and harmonic mean penalizes imbalanced communication.
Method recommendations:
| Method | Formula | Best for |
|---|---|---|
geometric |
General use, balanced | |
product |
Emphasizing strong bilateral signals | |
harmonic |
Penalizing imbalanced communication |
2. min_production and min_sensing: Noise
Filtering
# Compare different thresholds
thresholds <- c(0.05, 0.1, 0.2, 0.3)
n_interactions <- sapply(thresholds, function(t) {
obj_t <- computeCommunication(obj,
min_production = t, min_sensing = t,
n_permutations = 0, verbose = FALSE
)
sum(obj_t@communication_scores > 0)
})
barplot(n_interactions,
names.arg = thresholds,
xlab = "Threshold (min_production = min_sensing)",
ylab = "Number of Non-zero Interactions",
main = "Effect of Thresholds on Detected Interactions",
col = "#64B5F6"
)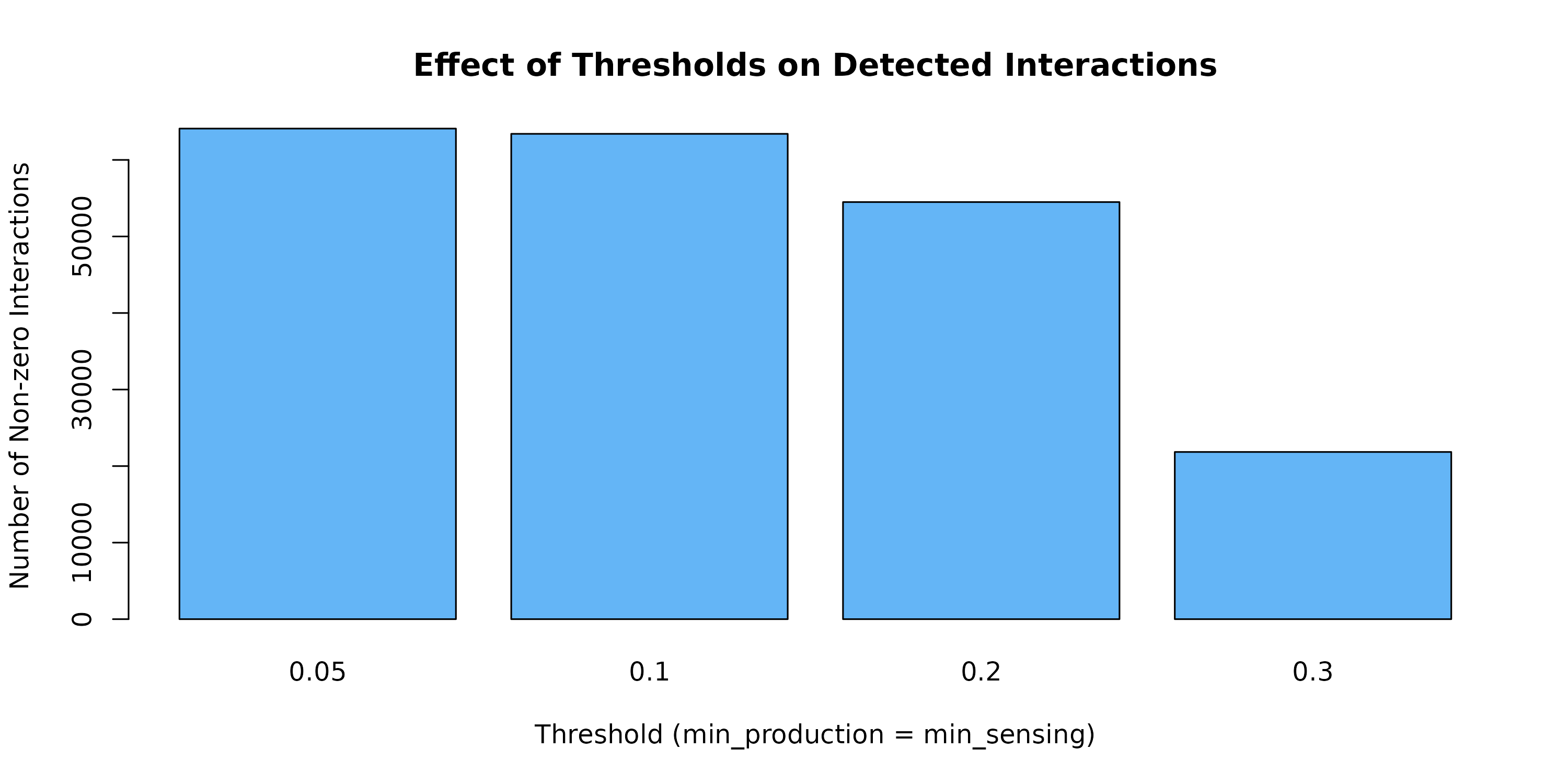
Figure 2: Effect of Expression Thresholds. Higher thresholds reduce the number of detected interactions by requiring stronger production and sensing signals. Choose based on desired stringency.
Guidelines: - 0.1: Default, moderate
filtering - 0.05: More permissive, may include noise -
0.2-0.3: More stringent, for focused analysis
3. population.size: Cell Type Abundance Weighting
When enabled, communication is weighted by cell type abundance:
# Compare with and without population size correction
obj_no_pop <- computeCommunication(obj,
population.size = FALSE,
n_permutations = 0, verbose = FALSE
)
obj_with_pop <- computeCommunication(obj,
population.size = TRUE,
n_permutations = 0, verbose = FALSE
)
# Total communication per cell type pair
total_no_pop <- apply(obj_no_pop@communication_scores, c(1, 2), sum)
total_with_pop <- apply(obj_with_pop@communication_scores, c(1, 2), sum)
cat("Without population size correction:\n")
#> Without population size correction:
print(round(total_no_pop[1:5, 1:5], 3))
#> receiver
#> sender Normal Epithelial T Plasma Normal Fibroblast CAF
#> Normal Epithelial 95.343 105.704 90.919 120.128 119.573
#> T 83.677 94.382 80.301 105.682 105.110
#> Plasma 83.028 93.692 80.557 104.482 103.873
#> Normal Fibroblast 91.996 102.945 87.495 114.784 114.247
#> CAF 91.263 101.869 86.869 113.753 113.273
cat("\nWith population size correction:\n")
#>
#> With population size correction:
print(round(total_with_pop[1:5, 1:5], 3))
#> receiver
#> sender Normal Epithelial T Plasma Normal Fibroblast CAF
#> Normal Epithelial 10.036 14.365 8.737 10.325 10.277
#> T 11.371 16.558 9.962 11.726 11.663
#> Plasma 7.978 11.623 7.066 8.197 8.150
#> Normal Fibroblast 7.907 11.422 6.865 8.055 8.017
#> CAF 7.844 11.303 6.816 7.983 7.949
# Show cell type sizes
cat("\nCell type counts:\n")
#>
#> Cell type counts:
print(table(crc_meta$cell_type))
#>
#> B CAF Endothelial Gliacyte
#> 150 200 100 20
#> Mast Monocyte Normal Epithelial Normal Fibroblast
#> 30 120 300 200
#> Normal Macrophage Pericyte Plasma SMC
#> 150 50 250 30
#> T TAM Tumor Epithelial
#> 500 150 600When to use population.size = TRUE: -
Tissue-level signaling analysis - When abundant cell types should have
more influence - Comparing communication strength across tissues
When to use population.size = FALSE: -
Cell type potential analysis (independent of abundance) - Comparing rare
vs common cell types fairly
Statistical Significance Testing
Permutation Test
# Run with permutation test
obj <- computeCommunication(obj, n_permutations = 100, verbose = TRUE)
#> | | | 0% | |= | 1% | |= | 2% | |== | 3% | |=== | 4% | |==== | 5% | |==== | 6% | |===== | 7% | |====== | 8% | |====== | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 13% | |========== | 14% | |========== | 15% | |=========== | 16% | |============ | 17% | |============= | 18% | |============= | 19% | |============== | 20% | |=============== | 21% | |=============== | 22% | |================ | 23% | |================= | 24% | |================== | 25% | |================== | 26% | |=================== | 27% | |==================== | 28% | |==================== | 29% | |===================== | 30% | |====================== | 31% | |====================== | 32% | |======================= | 33% | |======================== | 34% | |======================== | 35% | |========================= | 36% | |========================== | 37% | |=========================== | 38% | |=========================== | 39% | |============================ | 40% | |============================= | 41% | |============================= | 42% | |============================== | 43% | |=============================== | 44% | |================================ | 45% | |================================ | 46% | |================================= | 47% | |================================== | 48% | |================================== | 49% | |=================================== | 50% | |==================================== | 51% | |==================================== | 52% | |===================================== | 53% | |====================================== | 54% | |====================================== | 55% | |======================================= | 56% | |======================================== | 57% | |========================================= | 58% | |========================================= | 59% | |========================================== | 60% | |=========================================== | 61% | |=========================================== | 62% | |============================================ | 63% | |============================================= | 64% | |============================================== | 65% | |============================================== | 66% | |=============================================== | 67% | |================================================ | 68% | |================================================ | 69% | |================================================= | 70% | |================================================== | 71% | |================================================== | 72% | |=================================================== | 73% | |==================================================== | 74% | |==================================================== | 75% | |===================================================== | 76% | |====================================================== | 77% | |======================================================= | 78% | |======================================================= | 79% | |======================================================== | 80% | |========================================================= | 81% | |========================================================= | 82% | |========================================================== | 83% | |=========================================================== | 84% | |============================================================ | 85% | |============================================================ | 86% | |============================================================= | 87% | |============================================================== | 88% | |============================================================== | 89% | |=============================================================== | 90% | |================================================================ | 91% | |================================================================ | 92% | |================================================================= | 93% | |================================================================== | 94% | |================================================================== | 95% | |=================================================================== | 96% | |==================================================================== | 97% | |===================================================================== | 98% | |===================================================================== | 99% | |======================================================================| 100%
# Check p-value distribution
pvals <- as.vector(obj@communication_pvalues)
hist(pvals,
breaks = 50, main = "P-value Distribution",
xlab = "P-value", col = "#90CAF9", border = "white"
)
abline(v = 0.05, col = "red", lwd = 2, lty = 2)
text(0.1, par("usr")[4] * 0.9, "p = 0.05", col = "red")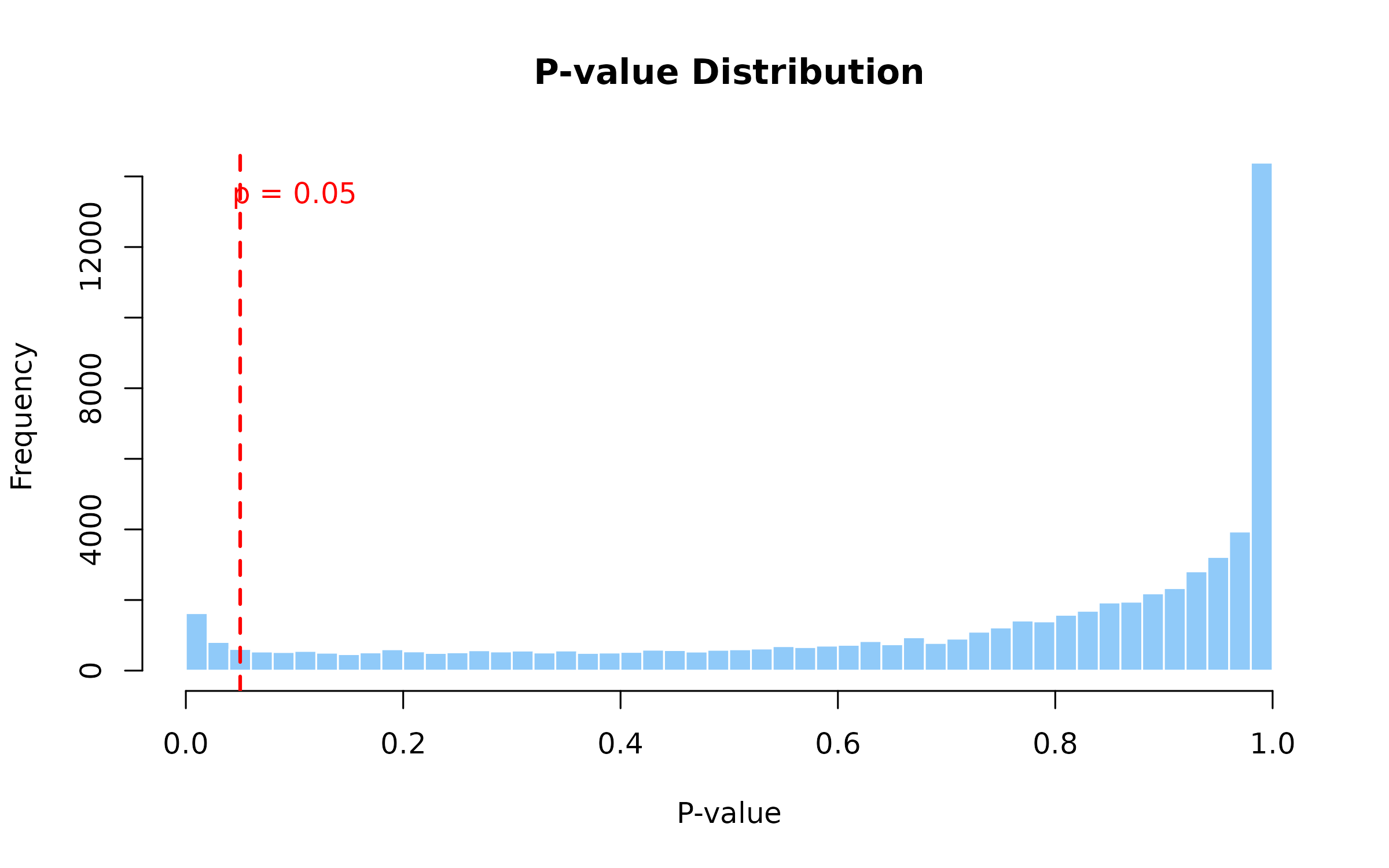
Figure 3: P-value Distribution from Permutation Test. Histogram of p-values from 100 permutations. A uniform distribution under the null is expected; enrichment near 0 indicates true signals. Red line marks the significance threshold (p=0.05).
Multiple Testing Correction
Different correction methods vary in stringency. With limited permutations (as in this tutorial), p-value correction may result in zero significant interactions:
# Different correction methods - show counts
obj_none <- filterSignificantInteractions(obj, adjust_method = "none")
obj_bh <- filterSignificantInteractions(obj, adjust_method = "BH")
cat("Significant interactions by correction method:\n")
#> Significant interactions by correction method:
cat(" No correction:", nrow(obj_none@significant_interactions), "\n")
#> No correction: 2754
cat(" BH correction:", nrow(obj_bh@significant_interactions), "\n")
#> BH correction: 0Note: In real analysis with sufficient permutations (>1000), BH correction is recommended. For demonstration purposes with limited permutations, we use no adjustment.
Filtering Significant Interactions
# For this tutorial, use no adjustment to demonstrate the workflow
# In real analysis, use adjust_method = "BH" with more permutations
obj <- filterSignificantInteractions(
obj,
pvalue_threshold = 0.05, # Significance level
adjust_method = "none", # Use "BH" for real analysis
min_score = 0 # Minimum communication score
)
# View results
cat("Total significant interactions:", nrow(obj@significant_interactions), "\n\n")
#> Total significant interactions: 2754
cat("Top interactions:\n")
#> Top interactions:
head(obj@significant_interactions[, c(
"sender", "receiver", "metabolite_name",
"communication_score", "pvalue_adjusted"
)])
#> sender receiver metabolite_name communication_score
#> 1 Monocyte Mast Epinephrine 0.9999641
#> 2 Normal Epithelial Plasma 5-Androstenediol 0.9992733
#> 3 Endothelial Mast Prostaglandin F1a 0.9903007
#> 4 Gliacyte B 12-Keto-leukotriene B4 0.9862472
#> 5 Plasma Plasma Estriol 0.9822572
#> 6 Normal Macrophage T Cholesterol sulfate 0.9768051
#> pvalue_adjusted
#> 1 0.00990099
#> 2 0.00990099
#> 3 0.00990099
#> 4 0.00990099
#> 5 0.00990099
#> 6 0.00990099Exploring Communication Results
Communication Score Structure
# The communication_scores slot is a 3D array
comm <- obj@communication_scores
cat("Dimensions: sender x receiver x metabolite\n")
#> Dimensions: sender x receiver x metabolite
cat(dim(comm), "\n")
#> 15 15 286
cat("\nCell types:", dimnames(comm)[[1]], "\n")
#>
#> Cell types: Normal Epithelial T Plasma Normal Fibroblast CAF Normal Macrophage TAM B Monocyte Endothelial Pericyte SMC Mast Gliacyte Tumor Epithelial
cat("\nMetabolites (first 10):", head(dimnames(comm)[[3]], 10), "\n")
#>
#> Metabolites (first 10): HMDB0000010 HMDB0000011 HMDB0000015 HMDB0000016 HMDB0000031 HMDB0000034 HMDB0000036 HMDB0000039 HMDB0000042 HMDB0000045Summarizing by Cell Type Pairs
# Different aggregation methods
sum_result <- summarizeCommunicationPairs(obj, aggregate_method = "sum")
mean_result <- summarizeCommunicationPairs(obj, aggregate_method = "mean")
count_result <- summarizeCommunicationPairs(obj, aggregate_method = "count")
cat("=== Sum of Communication Scores ===\n")
#> === Sum of Communication Scores ===
head(sum_result)
#> sender receiver communication_score
#> 128 Tumor Epithelial Normal Macrophage 52.63050
#> 199 Tumor Epithelial TAM 51.99507
#> 191 Normal Epithelial TAM 44.71782
#> 120 Normal Epithelial Normal Macrophage 43.56831
#> 106 Normal Epithelial Normal Fibroblast 40.51663
#> 30 Tumor Epithelial CAF 37.72378
cat("\n=== Mean Communication Score ===\n")
#>
#> === Mean Communication Score ===
head(mean_result)
#> sender receiver communication_score
#> 59 Endothelial Mast 0.9903007
#> 62 Monocyte Mast 0.8563538
#> 60 Gliacyte Mast 0.8530255
#> 66 Pericyte Mast 0.8407588
#> 132 Gliacyte Pericyte 0.8334445
#> 52 Pericyte Gliacyte 0.8238465
cat("\n=== Number of Significant Interactions ===\n")
#>
#> === Number of Significant Interactions ===
head(count_result)
#> sender receiver n_interactions
#> 128 Tumor Epithelial Normal Macrophage 82
#> 199 Tumor Epithelial TAM 80
#> 106 Normal Epithelial Normal Fibroblast 69
#> 30 Tumor Epithelial CAF 67
#> 113 Tumor Epithelial Normal Fibroblast 65
#> 191 Normal Epithelial TAM 63Get Communication Matrix
# Get aggregated communication matrix
comm_mat <- getCommunicationMatrix(obj, aggregate_method = "sum")
# Visualize
heatmap(comm_mat,
col = hcl.colors(50, "Reds"),
scale = "none",
main = "Cell-Cell Communication Strength"
)
Figure 7: Cell-Cell Communication Matrix. Aggregated communication strength between all cell type pairs. Rows represent senders, columns represent receivers. Clustering groups cell types with similar communication patterns.
Analyzing Specific Interactions
Top Communicating Pairs
sig <- obj@significant_interactions
# Top sender-receiver pairs
pair_counts <- aggregate(communication_score ~ sender + receiver, data = sig, FUN = length)
names(pair_counts)[3] <- "n_metabolites"
pair_counts <- pair_counts[order(-pair_counts$n_metabolites), ]
cat("Top communicating cell type pairs:\n")
#> Top communicating cell type pairs:
head(pair_counts, 10)
#> sender receiver n_metabolites
#> 128 Tumor Epithelial Normal Macrophage 82
#> 199 Tumor Epithelial TAM 80
#> 106 Normal Epithelial Normal Fibroblast 69
#> 30 Tumor Epithelial CAF 67
#> 113 Tumor Epithelial Normal Fibroblast 65
#> 191 Normal Epithelial TAM 63
#> 120 Normal Epithelial Normal Macrophage 61
#> 214 Tumor Epithelial Tumor Epithelial 58
#> 22 Normal Epithelial CAF 57
#> 123 Pericyte Normal Macrophage 47Top Metabolite Mediators
# Most frequent metabolites in significant interactions
met_counts <- table(sig$metabolite_name)
met_counts <- sort(met_counts, decreasing = TRUE)
cat("Top metabolite mediators:\n")
#> Top metabolite mediators:
head(met_counts, 15)
#>
#> 11beta-Hydroxyprogesterone 5-Androstenediol
#> 30 23
#> Ornithine 7beta-Hydroxycholesterol
#> 21 20
#> Adenosine monophosphate Cortisone
#> 19 19
#> Hydroxide Retinal
#> 19 18
#> L-Fucose Uridine 5'-diphosphate
#> 17 17
#> Cholesterol Cortisol
#> 16 16
#> Glycine Hyocholic acid
#> 16 16
#> L-Proline
#> 16Specific Pathway Focus
# Filter for specific metabolites (e.g., amino acids)
amino_acids <- c(
"L-Glutamic acid", "L-Glutamine", "L-Alanine", "Glycine",
"L-Serine", "L-Proline", "L-Aspartic acid"
)
aa_interactions <- sig[sig$metabolite_name %in% amino_acids, ]
cat("Amino acid-mediated interactions:", nrow(aa_interactions), "\n\n")
#> Amino acid-mediated interactions: 90
if (nrow(aa_interactions) > 0) {
# Summarize
aa_summary <- aggregate(communication_score ~ sender + receiver + metabolite_name,
data = aa_interactions, FUN = sum
)
aa_summary <- aa_summary[order(-aa_summary$communication_score), ]
head(aa_summary, 10)
}
#> sender receiver metabolite_name communication_score
#> 32 Gliacyte Tumor Epithelial L-Aspartic acid 0.8443290
#> 19 Normal Macrophage Normal Macrophage L-Alanine 0.8136641
#> 75 T Plasma L-Proline 0.8016842
#> 22 TAM Normal Macrophage L-Alanine 0.7858833
#> 3 Normal Macrophage Endothelial Glycine 0.7542903
#> 57 Normal Macrophage CAF L-Glutamine 0.7450304
#> 60 TAM CAF L-Glutamine 0.7300418
#> 74 Plasma Plasma L-Proline 0.7230354
#> 62 Normal Macrophage Normal Fibroblast L-Glutamine 0.7166993
#> 6 TAM Endothelial Glycine 0.7160865Advanced: Directional Analysis
Outgoing Communication (as Sender)
# Which cell types send the most signals?
outgoing <- aggregate(communication_score ~ sender, data = sig, FUN = sum)
outgoing <- outgoing[order(-outgoing$communication_score), ]
barplot(outgoing$communication_score,
names.arg = outgoing$sender,
las = 2, col = "#FF7043",
main = "Outgoing Communication Strength",
ylab = "Total Communication Score"
)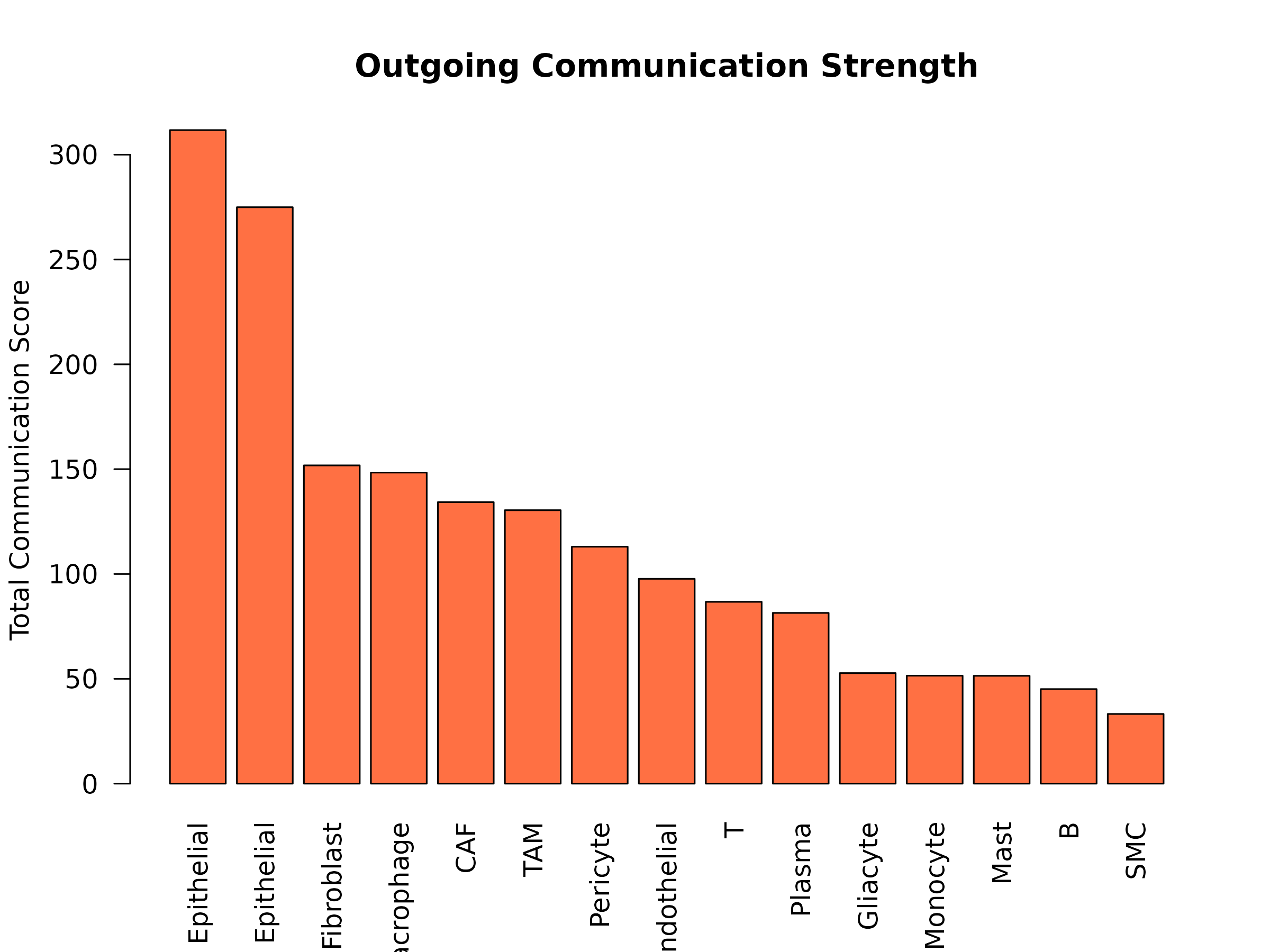
Figure 4: Outgoing Communication Strength. Total communication scores for each cell type acting as a sender. Higher values indicate cell types that produce more metabolites for intercellular signaling.
Incoming Communication (as Receiver)
# Which cell types receive the most signals?
incoming <- aggregate(communication_score ~ receiver, data = sig, FUN = sum)
incoming <- incoming[order(-incoming$communication_score), ]
barplot(incoming$communication_score,
names.arg = incoming$receiver,
las = 2, col = "#42A5F5",
main = "Incoming Communication Strength",
ylab = "Total Communication Score"
)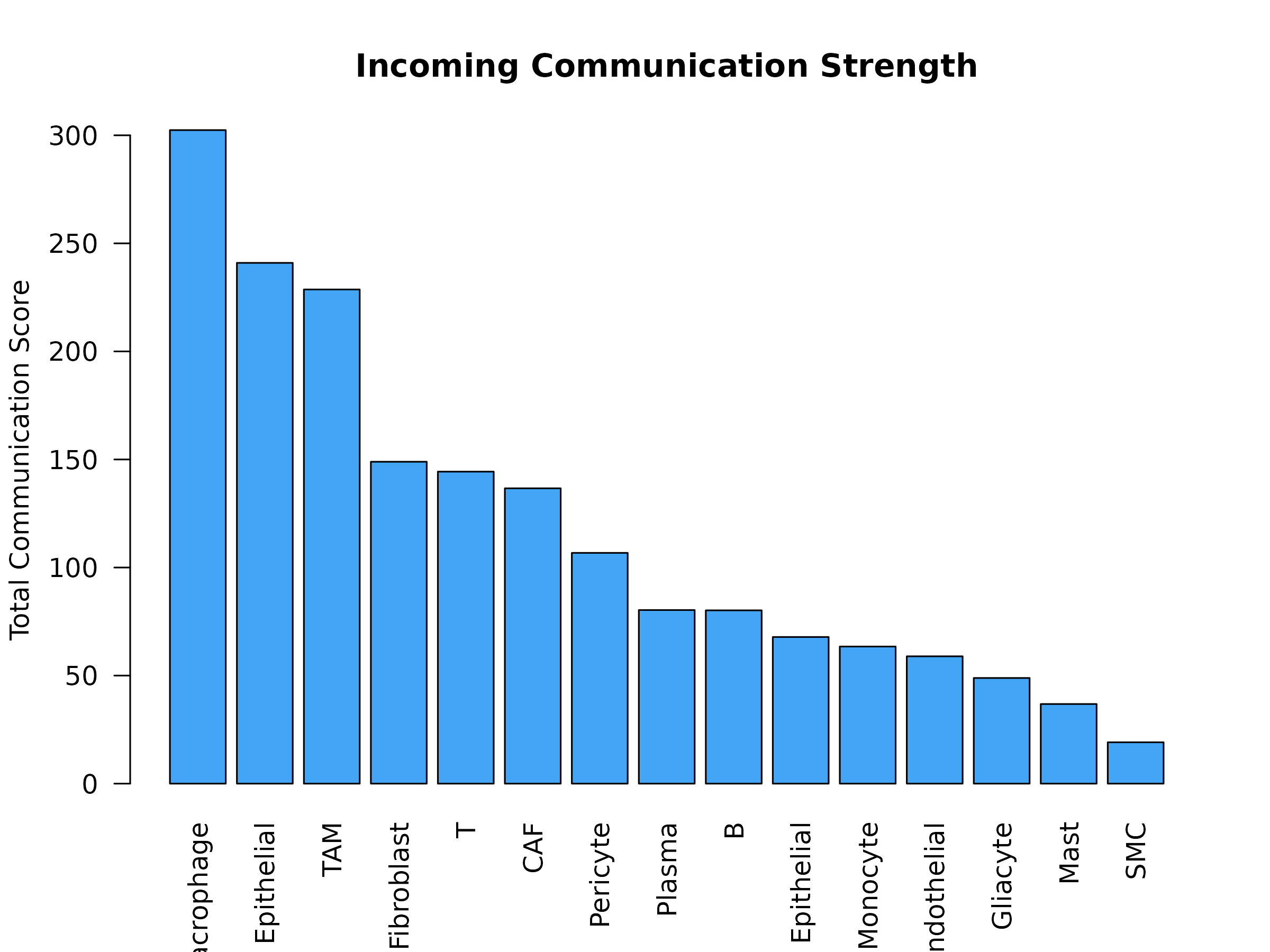
Figure 5: Incoming Communication Strength. Total communication scores for each cell type acting as a receiver. Higher values indicate cell types that sense/uptake more metabolites from other cells.
Net Communication Flow
# Net flow = Outgoing - Incoming
cell_types <- unique(c(sig$sender, sig$receiver))
net_flow <- sapply(cell_types, function(ct) {
out <- sum(sig$communication_score[sig$sender == ct])
inc <- sum(sig$communication_score[sig$receiver == ct])
out - inc
})
# Sort and plot
net_flow <- sort(net_flow, decreasing = TRUE)
cols <- ifelse(net_flow > 0, "#FF7043", "#42A5F5")
barplot(net_flow,
col = cols, las = 2,
main = "Net Communication Flow",
ylab = "Outgoing - Incoming"
)
abline(h = 0, lty = 2)
legend("topright",
legend = c("Net Sender", "Net Receiver"),
fill = c("#FF7043", "#42A5F5")
)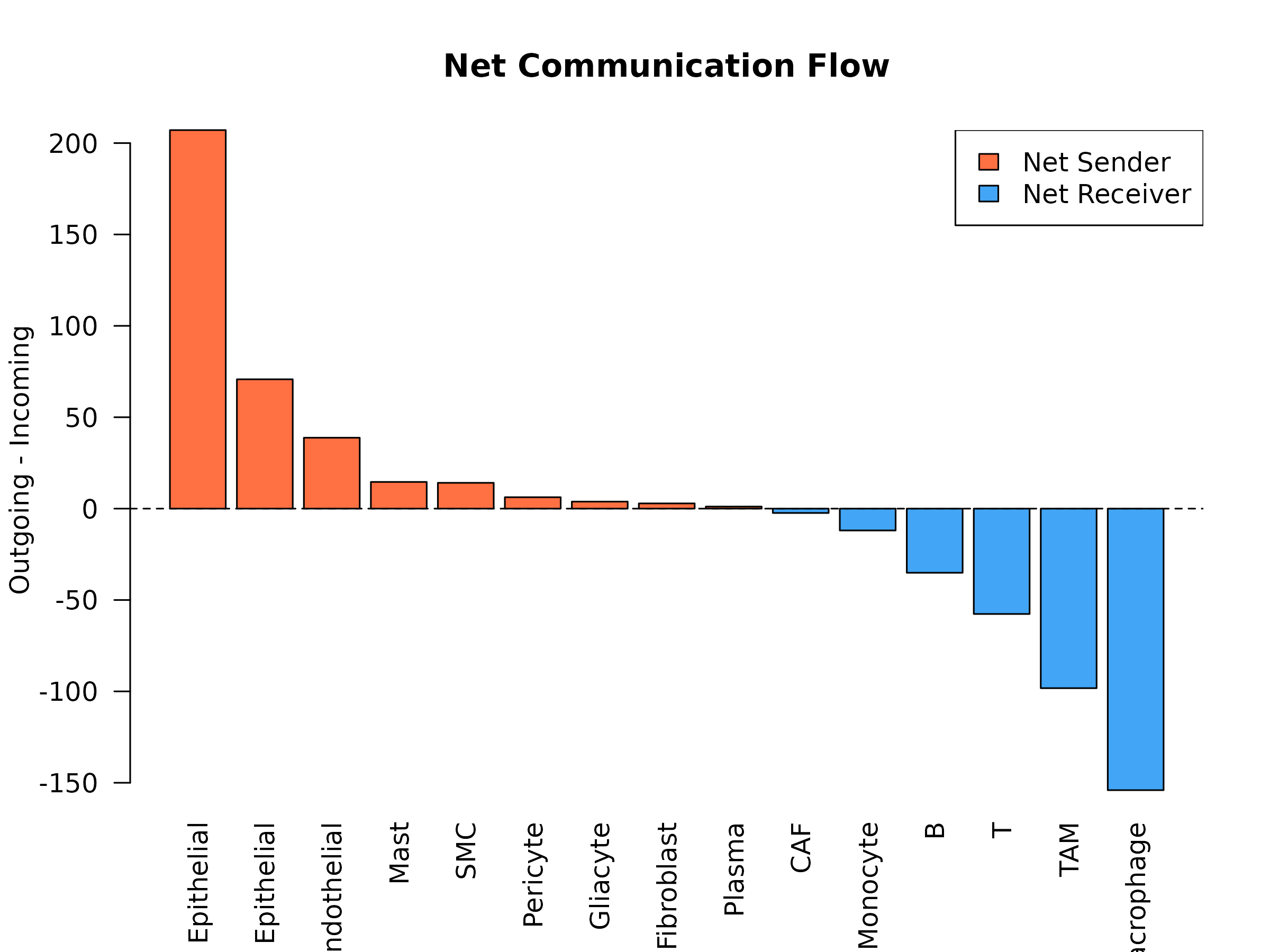
Figure 6: Net Communication Flow. Difference between outgoing and incoming communication for each cell type. Positive values (orange) indicate net senders; negative values (blue) indicate net receivers. This reveals which cell types drive vs receive metabolic signals in the tissue.
Comparing Conditions (If Available)
# If your data has conditions (e.g., tumor vs normal)
# You can run scMetaLink separately and compare
# Example workflow:
obj_tumor <- runScMetaLink(expr_tumor, meta_tumor, "cell_type")
obj_normal <- runScMetaLink(expr_normal, meta_normal, "cell_type")
# Compare communication strength
comm_tumor <- getCommunicationMatrix(obj_tumor)
comm_normal <- getCommunicationMatrix(obj_normal)
# Differential communication
diff_comm <- comm_tumor - comm_normalOutput Summary
| Output | Description | Access |
|---|---|---|
| Communication scores | 3D array (sender x receiver x metabolite) | obj@communication_scores |
| P-values | 3D array of permutation p-values | obj@communication_pvalues |
| Significant interactions | Filtered data.frame | obj@significant_interactions |
| Summary matrix | 2D aggregated matrix | getCommunicationMatrix(obj) |
| Pair summary | data.frame by cell type pairs | summarizeCommunicationPairs(obj) |
Next Steps
- Spatial Analysis: Spatial transcriptomics support
- Visualization: Create publication-ready figures
- Applications: Real-world analysis examples