Introduction
scMetaLink infers metabolite-mediated cell-cell communication from single-cell RNA-seq data. Unlike traditional ligand-receptor analysis, scMetaLink focuses on metabolites as signaling mediators, capturing a crucial but often overlooked layer of intercellular communication.
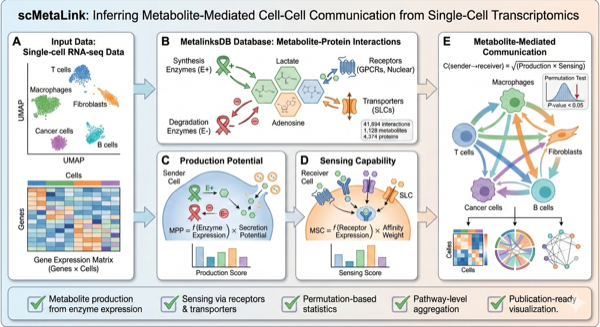
Figure 1: scMetaLink Workflow. Overview of the scMetaLink analysis pipeline.
Load Package and Example Data
library(scMetaLink)
library(Matrix)
# Load built-in colorectal cancer example data
data(crc_example)
# Check the data structure
cat("Expression matrix dimensions:", dim(crc_expr), "\n")
#> Expression matrix dimensions: 4210 2850
cat("Genes:", nrow(crc_expr), "| Cells:", ncol(crc_expr), "\n")
#> Genes: 4210 | Cells: 2850
cat("\nCell metadata:\n")
#>
#> Cell metadata:
head(crc_meta)
#> cell_type tumor_normal tissue_region
#> Cell_0001 Normal Epithelial Normal Normal Adjacent
#> Cell_0002 Normal Epithelial Normal Normal Adjacent
#> Cell_0003 Normal Epithelial Normal Normal Adjacent
#> Cell_0004 Normal Epithelial Normal Normal Adjacent
#> Cell_0005 Normal Epithelial Normal Normal Adjacent
#> Cell_0006 Normal Epithelial Normal Normal Adjacent
cat("\nCell type distribution:\n")
#>
#> Cell type distribution:
print(table(crc_meta$cell_type))
#>
#> B CAF Endothelial Gliacyte
#> 150 200 100 20
#> Mast Monocyte Normal Epithelial Normal Fibroblast
#> 30 120 300 200
#> Normal Macrophage Pericyte Plasma SMC
#> 150 50 250 30
#> T TAM Tumor Epithelial
#> 500 150 600The 5-Minute Workflow
Step 1: Create scMetaLink Object
# Create the analysis object
obj <- createScMetaLink(
expression_data = crc_expr,
cell_meta = crc_meta,
cell_type_column = "cell_type"
)
# Check the object
obj
#> scMetaLink Object
#> =================
#> Genes: 4210
#> Cells: 2850
#> Cell types: 15 (Normal Epithelial, T, Plasma, ...)Step 2: Infer Metabolite Production
Production potential reflects how capable each cell type is at synthesizing and secreting specific metabolites.
obj <- inferProduction(
obj,
method = "combined", # Use mean expression x proportion expressing
consider_degradation = TRUE, # Subtract degradation enzyme expression
consider_secretion = TRUE, # Weight by secretion potential
verbose = TRUE
)
# View top lactate producers
getTopProducers(obj, "L-Lactic acid", top_n = 5)
#> cell_type production_score rank
#> 1 Monocyte 0.9375074 1
#> 2 Gliacyte 0.5305421 2
#> 3 TAM 0.5021831 3
#> 4 Normal Macrophage 0.4832566 4
#> 5 Tumor Epithelial 0.3845369 5Step 3: Infer Metabolite Sensing
Sensing capability reflects how capable each cell type is at detecting and responding to specific metabolites.
obj <- inferSensing(
obj,
method = "combined",
weight_by_affinity = TRUE, # Weight by receptor-metabolite affinity
include_transporters = TRUE, # Include uptake transporters
verbose = TRUE
)
# View top glutamate sensors
getTopSensors(obj, "L-Glutamic acid", top_n = 5)
#> cell_type sensing_score rank
#> 1 Normal Macrophage 0.7624215 1
#> 2 TAM 0.6192208 2
#> 3 Tumor Epithelial 0.4362282 3
#> 4 Gliacyte 0.4255517 4
#> 5 Normal Fibroblast 0.4229089 5Step 4: Compute Communication
Communication scores quantify the potential signal flow from sender (producer) to receiver (sensor) cells.
obj <- computeCommunication(
obj,
method = "geometric", # sqrt(production x sensing)
min_production = 0.1, # Filter weak producers
min_sensing = 0.1, # Filter weak sensors
n_permutations = 100, # Permutation test for significance
verbose = TRUE
)
#> | | | 0% | |= | 1% | |= | 2% | |== | 3% | |=== | 4% | |==== | 5% | |==== | 6% | |===== | 7% | |====== | 8% | |====== | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 13% | |========== | 14% | |========== | 15% | |=========== | 16% | |============ | 17% | |============= | 18% | |============= | 19% | |============== | 20% | |=============== | 21% | |=============== | 22% | |================ | 23% | |================= | 24% | |================== | 25% | |================== | 26% | |=================== | 27% | |==================== | 28% | |==================== | 29% | |===================== | 30% | |====================== | 31% | |====================== | 32% | |======================= | 33% | |======================== | 34% | |======================== | 35% | |========================= | 36% | |========================== | 37% | |=========================== | 38% | |=========================== | 39% | |============================ | 40% | |============================= | 41% | |============================= | 42% | |============================== | 43% | |=============================== | 44% | |================================ | 45% | |================================ | 46% | |================================= | 47% | |================================== | 48% | |================================== | 49% | |=================================== | 50% | |==================================== | 51% | |==================================== | 52% | |===================================== | 53% | |====================================== | 54% | |====================================== | 55% | |======================================= | 56% | |======================================== | 57% | |========================================= | 58% | |========================================= | 59% | |========================================== | 60% | |=========================================== | 61% | |=========================================== | 62% | |============================================ | 63% | |============================================= | 64% | |============================================== | 65% | |============================================== | 66% | |=============================================== | 67% | |================================================ | 68% | |================================================ | 69% | |================================================= | 70% | |================================================== | 71% | |================================================== | 72% | |=================================================== | 73% | |==================================================== | 74% | |==================================================== | 75% | |===================================================== | 76% | |====================================================== | 77% | |======================================================= | 78% | |======================================================= | 79% | |======================================================== | 80% | |========================================================= | 81% | |========================================================= | 82% | |========================================================== | 83% | |=========================================================== | 84% | |============================================================ | 85% | |============================================================ | 86% | |============================================================= | 87% | |============================================================== | 88% | |============================================================== | 89% | |=============================================================== | 90% | |================================================================ | 91% | |================================================================ | 92% | |================================================================= | 93% | |================================================================== | 94% | |================================================================== | 95% | |=================================================================== | 96% | |==================================================================== | 97% | |===================================================================== | 98% | |===================================================================== | 99% | |======================================================================| 100%Step 5: Filter Significant Interactions
# For this quick tutorial with limited permutations, we use no adjustment
# In real analysis with more permutations, use adjust_method = "BH"
obj <- filterSignificantInteractions(
obj,
pvalue_threshold = 0.05,
adjust_method = "none" # Use "BH" for real analysis
)
# View top significant interactions
head(obj@significant_interactions, 10)
#> metabolite_id sender receiver communication_score pvalue
#> 1 HMDB0000068 Monocyte Mast 0.9999641 0.00990099
#> 2 HMDB0003818 Normal Epithelial Plasma 0.9992733 0.00990099
#> 3 HMDB0002685 Endothelial Mast 0.9903007 0.00990099
#> 4 HMDB0004234 Gliacyte B 0.9862472 0.00990099
#> 5 HMDB0000153 Plasma Plasma 0.9822572 0.00990099
#> 6 HMDB0000653 Normal Macrophage T 0.9768051 0.00990099
#> 7 HMDB0013078 Tumor Epithelial B 0.9747729 0.00990099
#> 8 HMDB0000190 Monocyte B 0.9679184 0.00990099
#> 9 HMDB0001139 Endothelial CAF 0.9659777 0.00990099
#> 10 HMDB0000335 Tumor Epithelial Plasma 0.9555982 0.00990099
#> pvalue_adjusted metabolite_name
#> 1 0.00990099 Epinephrine
#> 2 0.00990099 5-Androstenediol
#> 3 0.00990099 Prostaglandin F1a
#> 4 0.00990099 12-Keto-leukotriene B4
#> 5 0.00990099 Estriol
#> 6 0.00990099 Cholesterol sulfate
#> 7 0.00990099 Stearoylethanolamide
#> 8 0.00990099 L-Lactic acid
#> 9 0.00990099 Prostaglandin F2a
#> 10 0.00990099 16a-HydroxyestroneQuick Visualization
Communication Heatmap
The heatmap shows pairwise communication strength between cell types, aggregated across all significant metabolites.
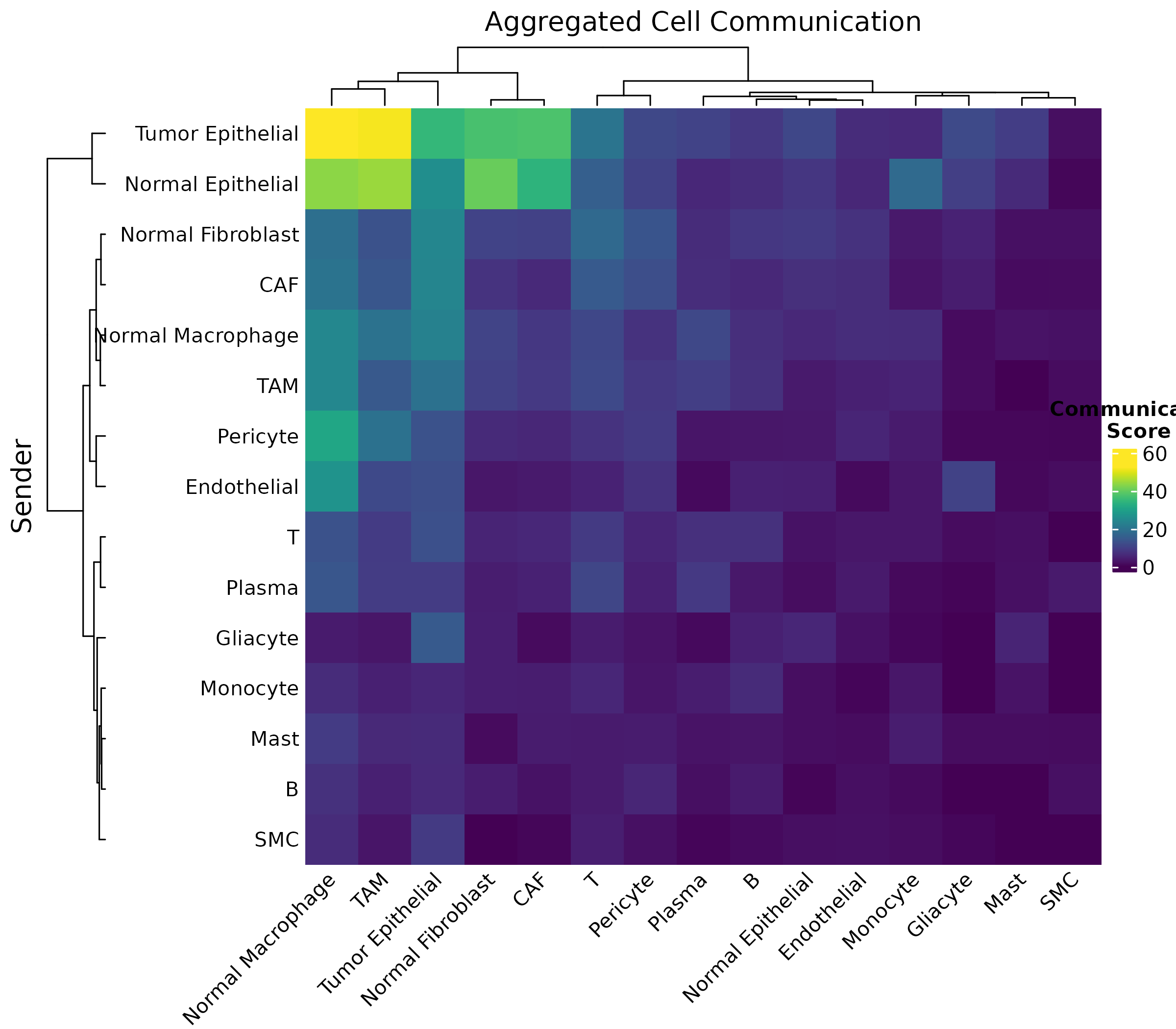
Figure 1: Communication Heatmap. Each cell shows the total communication score from sender (rows) to receiver (columns) cell types. Darker colors indicate stronger communication.
Chord Diagram
The chord diagram provides an intuitive view of communication flow between cell types.
plotCommunicationCircle(obj, top_n = 50)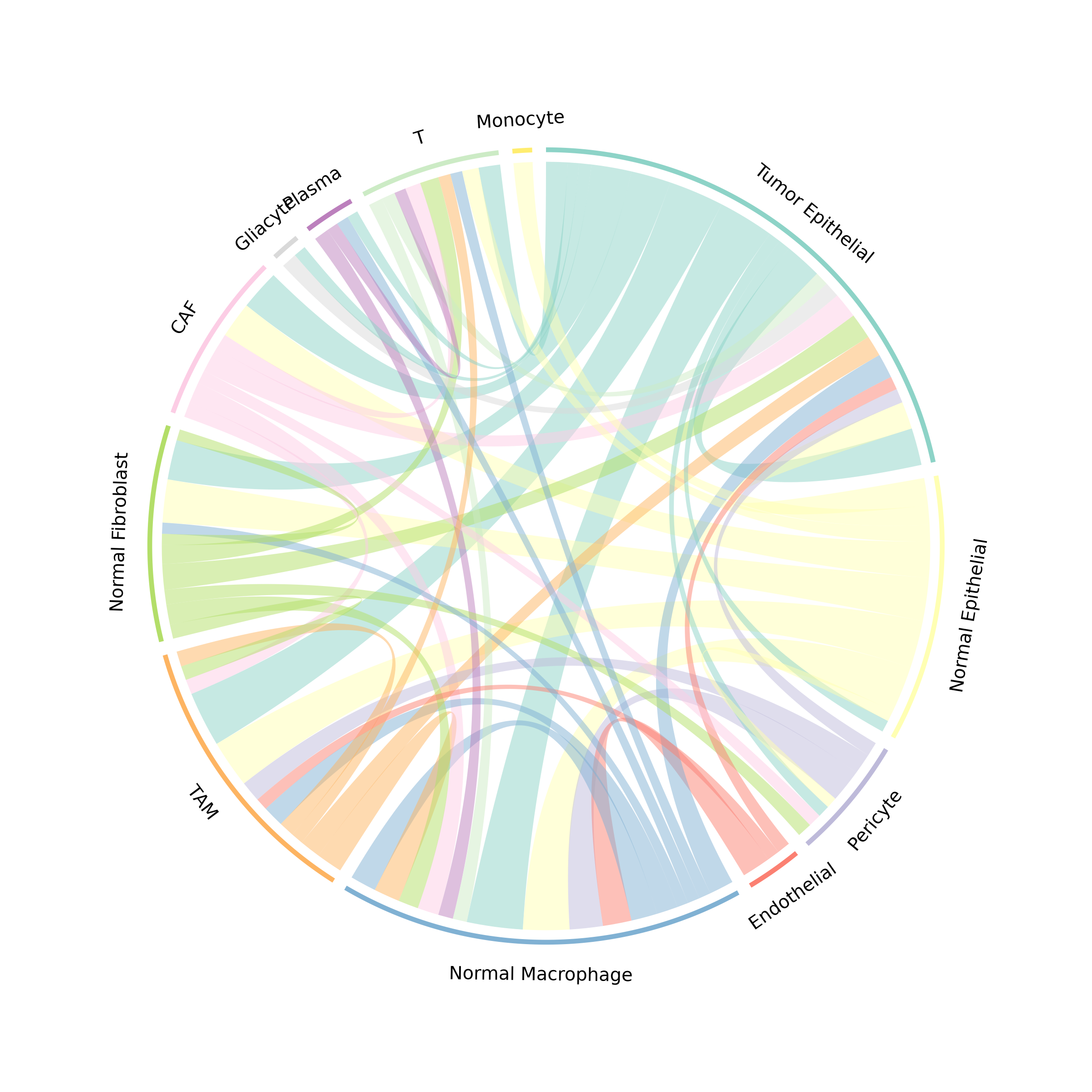
Figure 2: Communication Chord Diagram. Ribbons connect sender to receiver cell types, with width proportional to communication strength. Colors represent sender cell types.
One-Line Workflow
For convenience, you can run the entire analysis with a single function:
# Complete analysis in one line
obj <- runScMetaLink(
expression_data = crc_expr,
cell_meta = crc_meta,
cell_type_column = "cell_type",
n_permutations = 100
)Understanding the Output
Key Objects in the Result
| Slot | Description |
|---|---|
@production_scores |
Matrix: metabolites x cell types, production potential |
@sensing_scores |
Matrix: metabolites x cell types, sensing capability |
@communication_scores |
3D array: sender x receiver x metabolite |
@communication_pvalues |
3D array: permutation p-values |
@significant_interactions |
data.frame: filtered significant interactions |
Accessing Results
# Production scores (first 5 metabolites, first 5 cell types)
obj@production_scores[1:5, 1:5]
#> Normal Epithelial T Plasma Normal Fibroblast CAF
#> HMDB0000008 0.3844017 0.3124838 0.4670759 0.3124838 0.3124838
#> HMDB0000010 0.2708579 0.2465175 0.2391890 0.5269140 0.5625056
#> HMDB0000011 0.2804398 0.3580385 0.3628140 0.5658166 0.4273138
#> HMDB0000012 0.4768788 0.4473201 0.4021738 0.3852586 0.4177708
#> HMDB0000014 0.3271932 0.4098426 0.4522973 0.3559737 0.3869064
# Significant interactions summary
cat("Total significant interactions:", nrow(obj@significant_interactions), "\n")
#> Total significant interactions: 2754
cat("\nTop metabolites involved:\n")
#>
#> Top metabolites involved:
print(head(sort(table(obj@significant_interactions$metabolite_name), decreasing = TRUE), 10))
#>
#> 11beta-Hydroxyprogesterone 5-Androstenediol
#> 30 23
#> Ornithine 7beta-Hydroxycholesterol
#> 21 20
#> Adenosine monophosphate Cortisone
#> 19 19
#> Hydroxide Retinal
#> 19 18
#> L-Fucose Uridine 5'-diphosphate
#> 17 17Export Results
# Export all results to files
exportResults(obj, output_dir = "scMetaLink_results")Next Steps
- Theory & Methods: Understand the mathematical framework
- Production & Sensing: Deep dive into inference
- Communication Analysis: Advanced communication analysis
- Spatial Analysis: Spatial transcriptomics support
- Visualization: Complete visualization guide
- Applications: Real-world analysis examples
Session Info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] Matrix_1.7-4 scMetaLink_0.99.1
#>
#> loaded via a namespace (and not attached):
#> [1] viridis_0.6.5 sass_0.4.10 generics_0.1.4
#> [4] shape_1.4.6.1 lattice_0.22-7 digest_0.6.39
#> [7] magrittr_2.0.4 evaluate_1.0.5 grid_4.5.2
#> [10] RColorBrewer_1.1-3 iterators_1.0.14 circlize_0.4.17
#> [13] fastmap_1.2.0 foreach_1.5.2 doParallel_1.0.17
#> [16] jsonlite_2.0.0 GlobalOptions_0.1.3 gridExtra_2.3
#> [19] ComplexHeatmap_2.26.0 viridisLite_0.4.2 scales_1.4.0
#> [22] codetools_0.2-20 textshaping_1.0.4 jquerylib_0.1.4
#> [25] cli_3.6.5 rlang_1.1.7 crayon_1.5.3
#> [28] cachem_1.1.0 yaml_2.3.12 otel_0.2.0
#> [31] tools_4.5.2 parallel_4.5.2 dplyr_1.1.4
#> [34] colorspace_2.1-2 ggplot2_4.0.1 BiocGenerics_0.56.0
#> [37] GetoptLong_1.1.0 vctrs_0.7.1 R6_2.6.1
#> [40] png_0.1-8 stats4_4.5.2 matrixStats_1.5.0
#> [43] lifecycle_1.0.5 S4Vectors_0.48.0 IRanges_2.44.0
#> [46] fs_1.6.6 htmlwidgets_1.6.4 clue_0.3-66
#> [49] cluster_2.1.8.1 ragg_1.5.0 pkgconfig_2.0.3
#> [52] desc_1.4.3 pkgdown_2.2.0 pillar_1.11.1
#> [55] bslib_0.9.0 gtable_0.3.6 glue_1.8.0
#> [58] systemfonts_1.3.1 xfun_0.56 tibble_3.3.1
#> [61] tidyselect_1.2.1 knitr_1.51 farver_2.1.2
#> [64] rjson_0.2.23 htmltools_0.5.9 rmarkdown_2.30
#> [67] compiler_4.5.2 S7_0.2.1