Creates enrichment network visualizations showing connections between enriched terms and their associated genes. Terms and genes are both shown as nodes, with edges connecting terms to their genes.
Usage
EnrichNetwork(
data,
in_form = c("auto", "clusterProfiler", "clusterprofiler", "enrichr"),
split_by = NULL,
split_by_sep = "_",
top_term = 10,
metric = "p.adjust",
character_width = 50,
layout = "fr",
layoutadjust = TRUE,
adjscale = 60,
adjiter = 100,
blendmode = "blend",
labelsize = 5,
theme = "theme_ggforge",
theme_args = list(),
palette = "forge",
palcolor = NULL,
alpha = 1,
aspect.ratio = NULL,
legend.position = "right",
legend.direction = "vertical",
title = NULL,
subtitle = NULL,
xlab = NULL,
ylab = NULL,
seed = 8525,
combine = TRUE,
nrow = NULL,
ncol = NULL,
byrow = TRUE,
axes = NULL,
axis_titles = axes,
guides = NULL,
design = NULL,
...
)Arguments
- data
A data frame containing the data to be plotted. It should be in the format of clusterProfiler enrichment result, which includes the columns: ID, Description, GeneRatio, BgRatio, pvalue, p.adjust, qvalue, geneID and Count.
The
ID,qvalueandCountcolumns are optional.The
Descriptionis the description of the term.The
GeneRatiois the number of genes in the term divided by the total number of genes in the input list.The
BgRatiois the number of genes in the term divided by the total number of genes in the background list (all terms).The
Countcolumn, if given, should be the same as the first number in GeneRatio.
If you have enrichment results from multiple databases, you can combine them into one data frame and add a column (e.g. Database) to indicate the database. You can plot them in a single plot using the
split_byargument (e.g.split_by = "Database").- in_form
A character string specifying the input format. Either "auto", "clusterProfiler", "clusterprofiler" or "enrichr". The default is "auto", which will try to infer the input format.
- split_by
Column name(s) to split data into multiple plots
- split_by_sep
Separator when concatenating multiple split_by columns
- top_term
An integer specifying the number of top terms to show.
- metric
A character string specifying the metric to use for the size of the nodes. It is also used to order the terms when selected the top terms. Either "pvalue" or "p.adjust". The default is "p.adjust".
- character_width
The width of the characters used to wrap the keyword.
- layout
A character string specifying the layout of the graph. Either "circle", "tree", "grid" or other layout functions in
igraph.- layoutadjust
A logical value specifying whether to adjust the layout of the network.
- adjscale
A numeric value specifying the scale of the adjustment.
- adjiter
A numeric value specifying the number of iterations for the adjustment.
- blendmode
A character string specifying the blend mode of the colors. Either "blend", "average", "multiply" and "screen".
- labelsize
A numeric value specifying the size of the label.
- theme
Theme name (string) or theme function
- theme_args
List of arguments passed to theme function
- palette
Color palette name
- palcolor
Custom colors for palette
- alpha
Transparency level (0-1)
- aspect.ratio
Aspect ratio of plot panel
- legend.position
Legend position: "none", "left", "right", "bottom", "top"
- legend.direction
Legend direction: "horizontal" or "vertical"
- title
Plot title
- subtitle
Plot subtitle
- xlab
X-axis label
- ylab
Y-axis label
- seed
Random seed for reproducibility
- combine
Whether to combine split plots into one
- nrow
Number of rows when combining plots
- ncol
Number of columns when combining plots
- byrow
Fill combined plots by row
- axes
How to handle axes in combined plots ("keep", "collect", "collect_x", "collect_y")
- axis_titles
How to handle axis titles in combined plots
- guides
How to handle guides in combined plots ("collect", "keep", "auto")
- design
Custom layout design for combined plots
- ...
Additional arguments passed to atomic plotting functions.
See also
Other enrichment-plots:
EnrichMap(),
GSEAPlot(),
GSEASummaryPlot()
Examples
# \donttest{
data(enrich_example)
EnrichNetwork(enrich_example, top_term = 5)
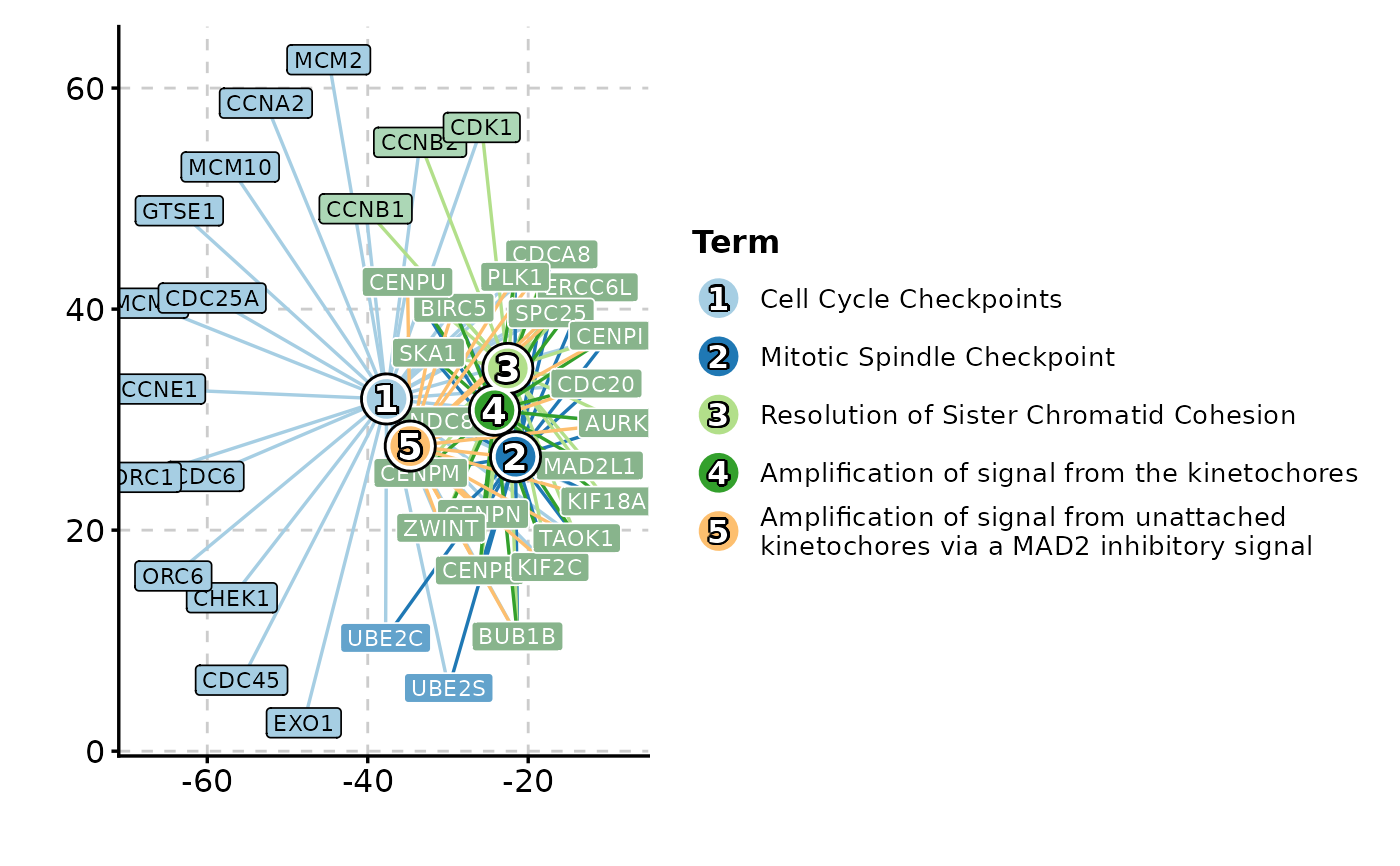 # }
# }
