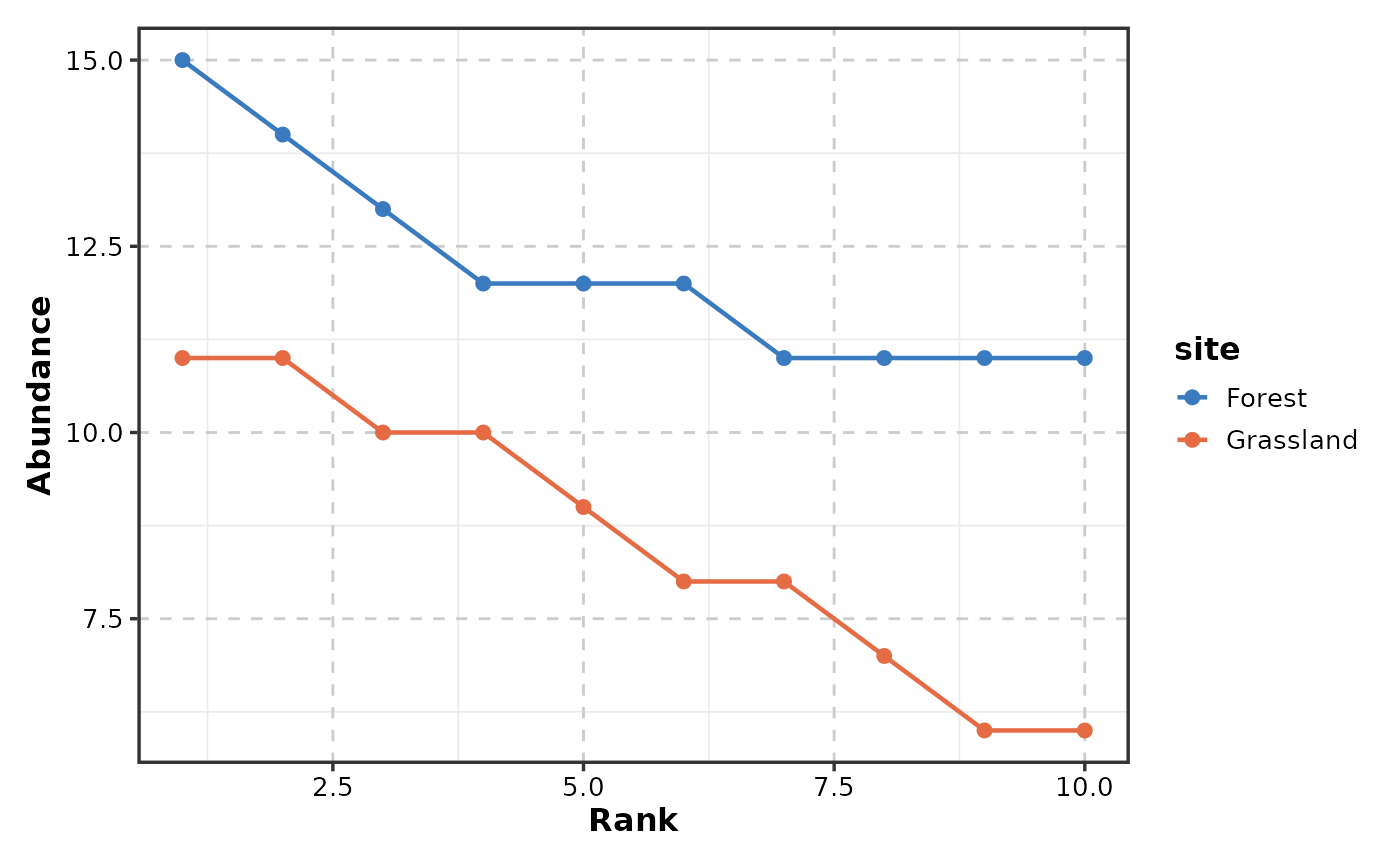
Creates rank-abundance curves (Whittaker plots) showing species abundance ranked from most to least abundant. Common in ecology for comparing community structure across sites or treatments.
Usage
RankAbundancePlot(
data,
species,
abundance,
group_by = NULL,
group_by_sep = "_",
group_name = NULL,
log_y = FALSE,
relative = FALSE,
pt_size = 2,
line_width = 0.8,
split_by = NULL,
split_by_sep = "_",
facet_by = NULL,
facet_scales = "fixed",
facet_nrow = NULL,
facet_ncol = NULL,
facet_byrow = TRUE,
theme = "theme_ggforge_grid",
theme_args = list(),
palette = "forge",
palcolor = NULL,
alpha = 1,
legend.position = "right",
legend.direction = "vertical",
title = NULL,
subtitle = NULL,
xlab = NULL,
ylab = NULL,
seed = 8525,
combine = TRUE,
nrow = NULL,
ncol = NULL,
byrow = TRUE,
axes = NULL,
axis_titles = NULL,
guides = NULL,
design = NULL,
...
)Arguments
- data
A data frame containing the data to plot
- species
Column for species/taxon names.
- abundance
Column for abundance values.
- group_by
Column for grouping (different communities).
- group_by_sep
Separator for multiple group_by columns.
- group_name
Legend title.
- log_y
Whether to log-transform the y-axis.
- relative
Whether to convert abundance to relative (proportion).
- pt_size
Point size.
- line_width
Line width.
- split_by
Column name(s) to split data into multiple plots
- split_by_sep
Separator when concatenating multiple split_by columns
- facet_by
Column name(s) for faceting the plot
- facet_scales
Scales for facets: "fixed", "free", "free_x", "free_y"
- facet_nrow
Number of rows in facet layout
- facet_ncol
Number of columns in facet layout
- facet_byrow
Fill facets by row (TRUE) or column (FALSE)
- theme
Theme name (string) or theme function
- theme_args
List of arguments passed to theme function
- palette
Color palette name
- palcolor
Custom colors for palette
- alpha
Transparency level (0-1)
- legend.position
Legend position: "none", "left", "right", "bottom", "top"
- legend.direction
Legend direction: "horizontal" or "vertical"
- title
Plot title
- subtitle
Plot subtitle
- xlab
X-axis label
- ylab
Y-axis label
- seed
Random seed for reproducibility
- combine
Whether to combine split plots into one
- nrow
Number of rows when combining plots
- ncol
Number of columns when combining plots
- byrow
Fill combined plots by row
- axes
How to handle axes in combined plots ("keep", "collect", "collect_x", "collect_y")
- axis_titles
How to handle axis titles in combined plots
- guides
How to handle guides in combined plots ("collect", "keep", "auto")
- design
Custom layout design for combined plots
- ...
Additional arguments passed to atomic plotting functions.
See also
Other ecology-evolution-plots:
OrdinationPlot(),
PhyloTreePlot()