Getting Started with darwin
Zaoqu Liu
2026-01-25
Source:vignettes/darwin-intro.Rmd
darwin-intro.RmdIntroduction
darwin is an R package for automatic marker gene selection using multi-objective evolutionary optimization. The package implements the NSGA-II algorithm to identify Pareto-optimal gene subsets for bulk RNA-seq deconvolution.
Why darwin?
Traditional marker gene selection often relies on single-objective criteria, which may lead to suboptimal solutions. darwin addresses this by:
- Multi-objective optimization: Simultaneously balancing multiple criteria
- Pareto optimality: Providing a diverse set of trade-off solutions
- Automated selection: Reducing manual intervention in gene selection
Installation
# From R-universe (recommended)
install.packages("darwin", repos = "https://zaoqu-liu.r-universe.dev")
# From GitHub
remotes::install_github("Zaoqu-Liu/darwin")Quick Start
Prepare Reference Data
darwin requires a reference expression matrix where rows are cell types and columns are genes.
set.seed(42)
# Simulate reference data: 5 cell types × 200 genes
n_celltypes <- 5
n_genes <- 200
reference <- matrix(
abs(rnorm(n_celltypes * n_genes, mean = 2, sd = 1)),
nrow = n_celltypes,
ncol = n_genes
)
rownames(reference) <- paste0("CellType", 1:n_celltypes)
colnames(reference) <- paste0("Gene", 1:n_genes)
# Add cell-type specific marker genes
for (i in 1:n_celltypes) {
marker_start <- (i - 1) * 10 + 1
marker_end <- i * 10
reference[i, marker_start:marker_end] <- reference[i, marker_start:marker_end] + 5
}
print(dim(reference))
#> [1] 5 200Initialize darwin
dw <- darwin(reference)Visualize Pareto Front
dw$plot()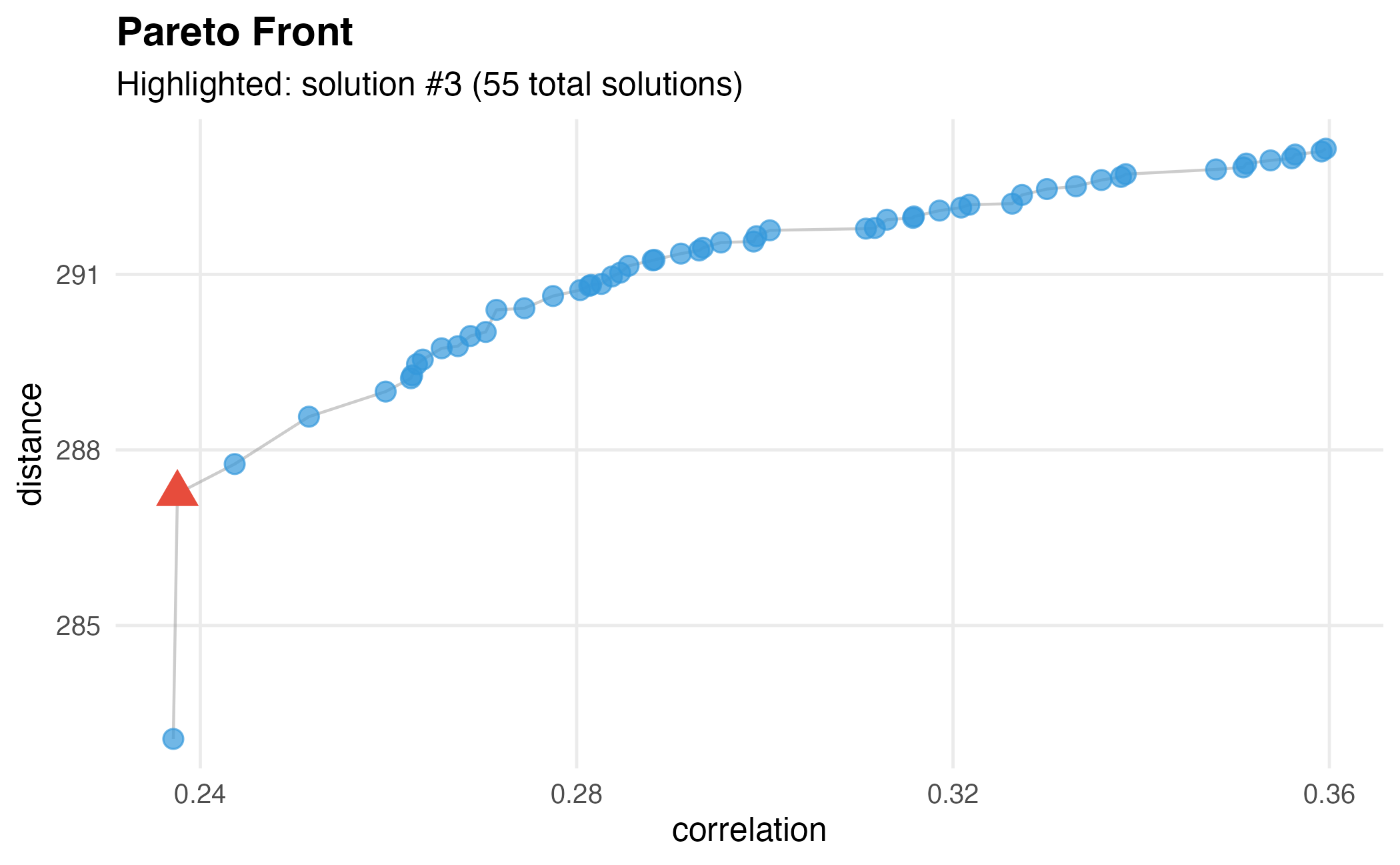
Pareto front showing the trade-off between correlation and distance objectives.
Select Optimal Solution
# Select using weighted criteria
dw$select(weights = c(-1, 1))
# Get selected genes
genes <- dw$get_genes()
cat("Number of selected genes:", length(genes), "\n")
#> Number of selected genes: 191
cat("First 10 genes:", paste(head(genes, 10), collapse = ", "), "\n")
#> First 10 genes: Gene1, Gene2, Gene3, Gene4, Gene5, Gene6, Gene7, Gene8, Gene9, Gene10View Fitness Values
fitness <- dw$get_fitness()
head(fitness)
#> correlation distance
#> 1 0.2371333 283.0623
#> 2 0.3596163 293.1472
#> 3 0.2375669 287.2533
#> 4 0.2515467 288.5683
#> 5 0.2597023 288.9972
#> 6 0.2436569 287.7592Basic Deconvolution
# Create mock bulk data
bulk <- matrix(abs(rnorm(3 * n_genes)), nrow = 3, ncol = n_genes)
colnames(bulk) <- colnames(reference)
rownames(bulk) <- paste0("Sample", 1:3)
# Perform deconvolution
result <- dw$deconvolve(bulk, method = "nnls")
# View estimated proportions
print(round(result$proportions, 3))
#> CellType1 CellType2 CellType3 CellType4 CellType5
#> Sample1 0.210 0.171 0.275 0.148 0.197
#> Sample2 0.078 0.131 0.299 0.233 0.261
#> Sample3 0.189 0.178 0.206 0.167 0.260Session Info
sessionInfo()
#> R version 4.4.0 (2024-04-24)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] C
#>
#> time zone: Asia/Shanghai
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] darwin_1.0.0
#>
#> loaded via a namespace (and not attached):
#> [1] sass_0.4.10 future_1.69.0 generics_0.1.4
#> [4] lattice_0.22-7 listenv_0.10.0 digest_0.6.39
#> [7] magrittr_2.0.4 evaluate_1.0.5 grid_4.4.0
#> [10] RColorBrewer_1.1-3 fastmap_1.2.0 jsonlite_2.0.0
#> [13] Matrix_1.7-4 scales_1.4.0 codetools_0.2-20
#> [16] textshaping_1.0.4 jquerylib_0.1.4 cli_3.6.5
#> [19] rlang_1.1.7 parallelly_1.46.1 future.apply_1.20.1
#> [22] withr_3.0.2 cachem_1.1.0 yaml_2.3.12
#> [25] otel_0.2.0 tools_4.4.0 parallel_4.4.0
#> [28] dplyr_1.1.4 ggplot2_4.0.1 globals_0.18.0
#> [31] vctrs_0.7.1 R6_2.6.1 lifecycle_1.0.5
#> [34] fs_1.6.6 htmlwidgets_1.6.4 ragg_1.5.0
#> [37] pkgconfig_2.0.3 desc_1.4.3 pkgdown_2.1.3
#> [40] pillar_1.11.1 bslib_0.9.0 gtable_0.3.6
#> [43] glue_1.8.0 Rcpp_1.1.1 systemfonts_1.3.1
#> [46] xfun_0.56 tibble_3.3.1 tidyselect_1.2.1
#> [49] knitr_1.51 dichromat_2.0-0.1 farver_2.1.2
#> [52] htmltools_0.5.9 rmarkdown_2.30 labeling_0.4.3
#> [55] compiler_4.4.0 S7_0.2.1 nnls_1.6