Create violin plots with optional grouping, faceting, and statistical comparisons. Violin plots combine box plots with kernel density estimation to show the distribution shape of continuous data.
Usage
ViolinPlot(
data,
x,
x_sep = "_",
y = NULL,
in_form = c("long", "wide"),
split_by = NULL,
split_by_sep = "_",
symnum_args = NULL,
sort_x = NULL,
flip = FALSE,
keep_empty = FALSE,
group_by = NULL,
group_by_sep = "_",
group_name = NULL,
paired_by = NULL,
x_text_angle = NULL,
step_increase = 0.1,
fill_mode = ifelse(!is.null(group_by), "dodge", "x"),
fill_reverse = FALSE,
theme = "theme_ggforge",
theme_args = list(),
palette = "forge",
palcolor = NULL,
alpha = 1,
aspect.ratio = NULL,
legend.position = "right",
legend.direction = "vertical",
add_point = FALSE,
pt_color = "grey30",
pt_size = NULL,
pt_alpha = 1,
jitter_width = NULL,
jitter_height = 0,
add_beeswarm = FALSE,
beeswarm_method = "swarm",
beeswarm_cex = 1,
beeswarm_priority = "ascending",
beeswarm_dodge = 0.9,
stack = FALSE,
y_max = NULL,
y_min = NULL,
add_box = FALSE,
box_color = "black",
box_width = 0.1,
box_ptsize = 2.5,
add_trend = FALSE,
trend_color = NULL,
trend_linewidth = 1,
trend_ptsize = 2,
add_stat = NULL,
stat_name = NULL,
stat_color = "black",
stat_size = 1,
stat_stroke = 1,
stat_shape = 25,
add_bg = FALSE,
bg_palette = "stripe",
bg_palcolor = NULL,
bg_alpha = 0.2,
add_line = NULL,
line_color = "red2",
line_width = 0.6,
line_type = 2,
highlight = NULL,
highlight_color = "red2",
highlight_size = 1,
highlight_alpha = 1,
comparisons = NULL,
ref_group = NULL,
pairwise_method = "wilcox.test",
multiplegroup_comparisons = FALSE,
multiple_method = "kruskal.test",
sig_label = "p.format",
sig_labelsize = 3.5,
hide_ns = FALSE,
facet_by = NULL,
facet_scales = "fixed",
facet_ncol = NULL,
facet_nrow = NULL,
facet_byrow = TRUE,
title = NULL,
subtitle = NULL,
xlab = NULL,
ylab = NULL,
seed = 8525,
combine = TRUE,
nrow = NULL,
ncol = NULL,
byrow = TRUE,
axes = NULL,
axis_titles = axes,
guides = NULL,
...
)Arguments
- data
A data frame containing the data to plot
- x
Column for x-axis (discrete). Can be a single column name or multiple columns that will be concatenated.
- x_sep
Separator for concatenating multiple x columns.
- y
Column for y-axis (numeric). The response variable.
- in_form
Input data form: "long" (default) or "wide"
- split_by
Column name(s) to split data into multiple plots
- split_by_sep
Separator when concatenating multiple split_by columns
- symnum_args
Symbolic number coding arguments for significance
- sort_x
Expression string to sort x-axis values, or NULL for no sorting. Legacy values supported: "none", "mean_asc", "mean_desc", "mean", "median_asc", "median_desc", "median". Custom expressions also accepted (e.g., "mean(value)", "-median(value)").
- flip
Logical; flip coordinates to create horizontal plots
- keep_empty
Logical; keep empty factor levels on x-axis
- group_by
Column for grouping (creates dodged/side-by-side plots)
- group_by_sep
Separator when concatenating multiple group_by columns
- group_name
Legend name for groups
- paired_by
Column identifying paired observations (for paired tests)
- x_text_angle
Angle for x-axis text labels
- step_increase
Step increase for comparison brackets
- fill_mode
Fill coloring mode: "dodge", "x", "mean", or "median"
- fill_reverse
Logical; reverse gradient fills
- theme
Theme name (string) or theme function
- theme_args
List of arguments passed to theme function
- palette
Color palette name
- palcolor
Custom colors for palette
- alpha
Transparency level (0-1)
- aspect.ratio
Aspect ratio of plot panel
- legend.position
Legend position: "none", "left", "right", "bottom", "top"
- legend.direction
Legend direction: "horizontal" or "vertical"
- add_point
Logical; add jittered data points
- pt_color
Point color (default: "grey30")
- pt_size
Point size (auto-calculated if NULL)
- pt_alpha
Point transparency (0-1)
- jitter_width
Jitter width for points
- jitter_height
Jitter height for points
- add_beeswarm
Logical; use beeswarm layout instead of jitter for points. Requires the ggbeeswarm package. Not compatible with paired_by.
- beeswarm_method
Beeswarm method: "swarm", "compactswarm", "hex", "square", or "center"
- beeswarm_cex
Scaling for point spacing in beeswarm (default: 1)
- beeswarm_priority
Point layout priority: "ascending", "descending", "density", or "random"
- beeswarm_dodge
Dodge width for grouped beeswarm points (default: 0.9)
- stack
Logical; stack facets vertically/horizontally
- y_max
Y-axis maximum (numeric or "qXX" for quantile)
- y_min
Y-axis minimum (numeric or "qXX" for quantile)
- add_box
Logical; add box overlay (violin only)
- box_color
Box overlay color
- box_width
Box overlay width
- box_ptsize
Box median point size
- add_trend
Logical; add trend line connecting medians
- trend_color
Trend line color
- trend_linewidth
Trend line width
- trend_ptsize
Trend point size
- add_stat
Function to add stat summary (e.g., mean)
- stat_name
Stat legend name
- stat_color
Stat point color
- stat_size
Stat point size
- stat_stroke
Stat point stroke width
- stat_shape
Stat point shape
- add_bg
Logical; add alternating background
- bg_palette
Background color palette
- bg_palcolor
Background custom colors
- bg_alpha
Background transparency
- add_line
Numeric; add horizontal reference line at this value
- line_color
Reference line color
- line_width
Reference line width
- line_type
Reference line type
- highlight
Points to highlight (logical, indices, or expression)
- highlight_color
Highlight color
- highlight_size
Highlight size
- highlight_alpha
Highlight transparency
- comparisons
Pairwise comparisons (list of pairs or TRUE for all)
- ref_group
Reference group for comparisons
- pairwise_method
Statistical method for pairwise comparisons
- multiplegroup_comparisons
Logical; perform multiple group comparisons
- multiple_method
Statistical method for multiple comparisons
- sig_label
Significance label format: "p.format" or "p.signif"
- sig_labelsize
Significance label font size
- hide_ns
Logical; hide non-significant comparisons
- facet_by
Column name(s) for faceting the plot
- facet_scales
Scales for facets: "fixed", "free", "free_x", "free_y"
- facet_ncol
Number of columns in facet layout
- facet_nrow
Number of rows in facet layout
- facet_byrow
Fill facets by row (TRUE) or column (FALSE)
- title
Plot title
- subtitle
Plot subtitle
- xlab
X-axis label
- ylab
Y-axis label
- seed
Random seed for reproducibility
- combine
Whether to combine split plots into one
- nrow
Number of rows when combining plots
- ncol
Number of columns when combining plots
- byrow
Fill combined plots by row
- axes
How to handle axes in combined plots ("keep", "collect", "collect_x", "collect_y")
- axis_titles
How to handle axis titles in combined plots
- guides
How to handle guides in combined plots ("collect", "keep", "auto")
- ...
Additional arguments passed to atomic plotting functions.
See also
Examples
# \donttest{
# ============================================================
# Basic Examples
# ============================================================
# Create sample data with different distributions
set.seed(456)
data <- data.frame(
category = rep(c("Normal", "Bimodal", "Skewed", "Uniform"), each = 100),
value = c(
rnorm(100, 50, 10), # Normal
c(rnorm(50, 30, 5), rnorm(50, 70, 5)), # Bimodal
rexp(100, 0.1), # Skewed
runif(100, 20, 80) # Uniform
),
group = rep(c("A", "B"), 200)
)
# Simple violin plot
ViolinPlot(data, x = "category", y = "value")
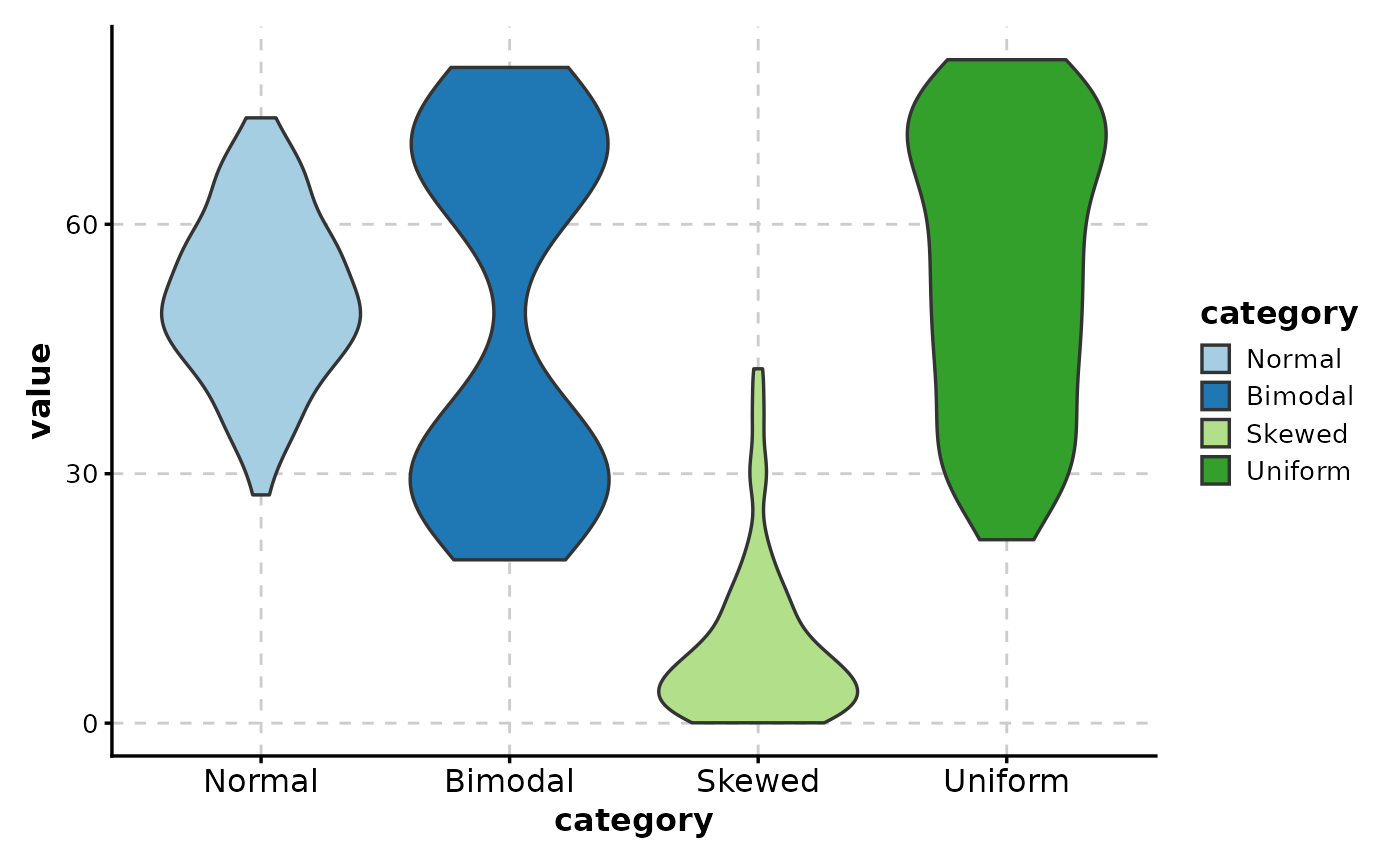 # With title and labels
ViolinPlot(data,
x = "category", y = "value",
title = "Distribution Shapes Comparison",
xlab = "Distribution Type",
ylab = "Value"
)
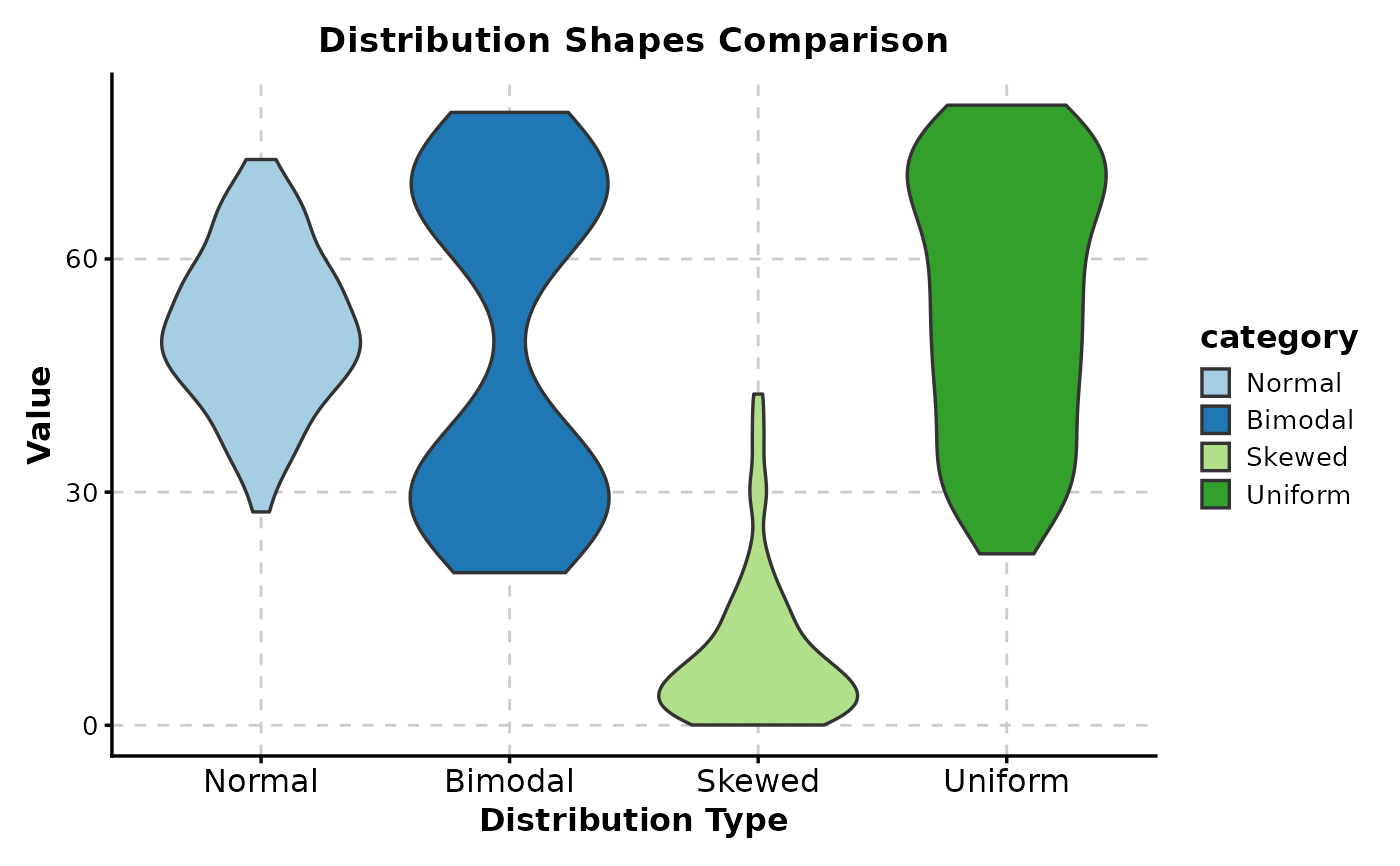
# With title and labels
ViolinPlot(data,
x = "category", y = "value",
title = "Distribution Shapes Comparison",
xlab = "Distribution Type",
ylab = "Value"
)
 # ============================================================
# Violin with Box Plot Overlay
# ============================================================
# Add box plot inside violin (shows quartiles)
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE
)
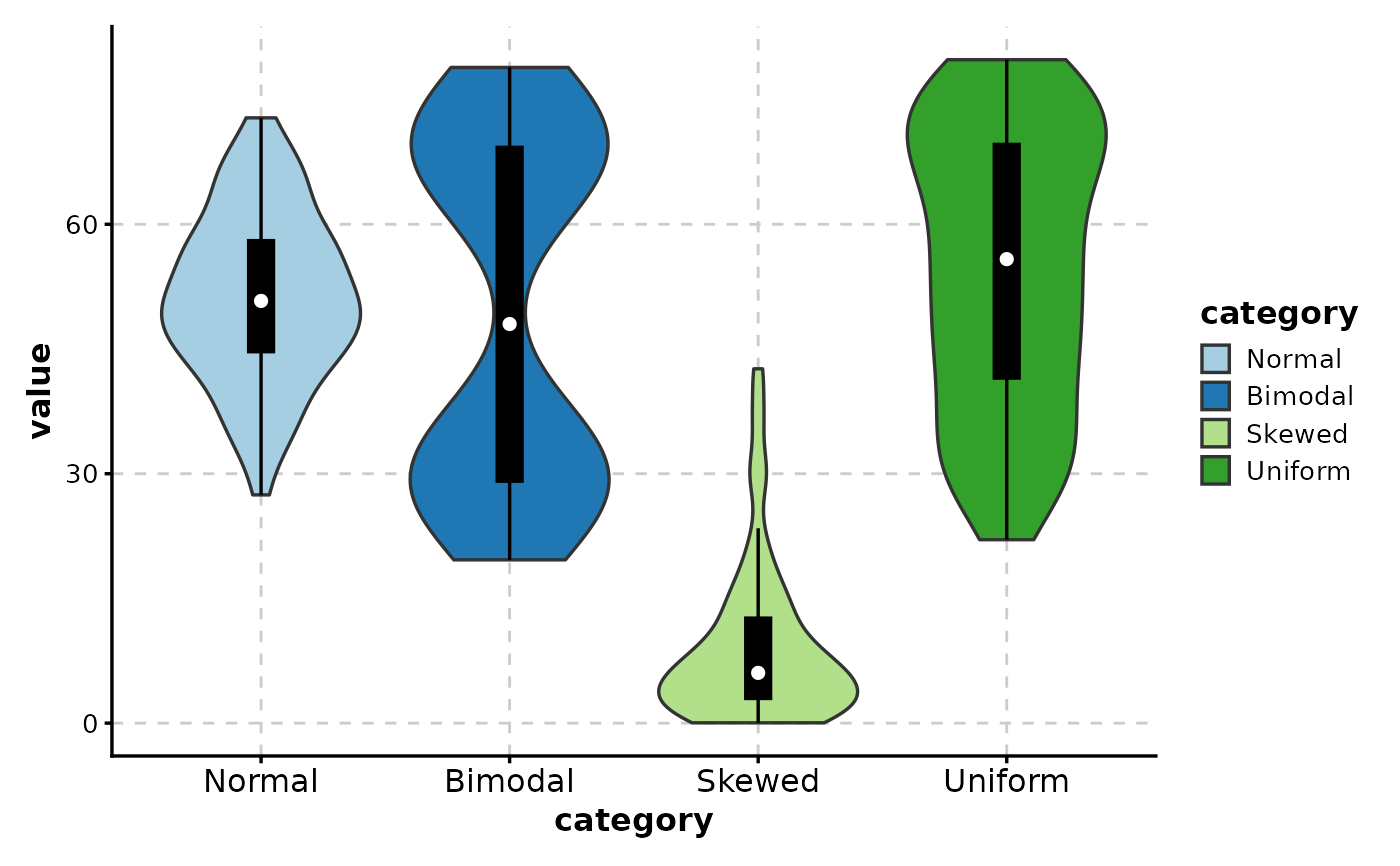
# ============================================================
# Violin with Box Plot Overlay
# ============================================================
# Add box plot inside violin (shows quartiles)
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE
)
 # Customize box overlay
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE,
box_color = "white",
box_width = 0.15,
box_ptsize = 3
)
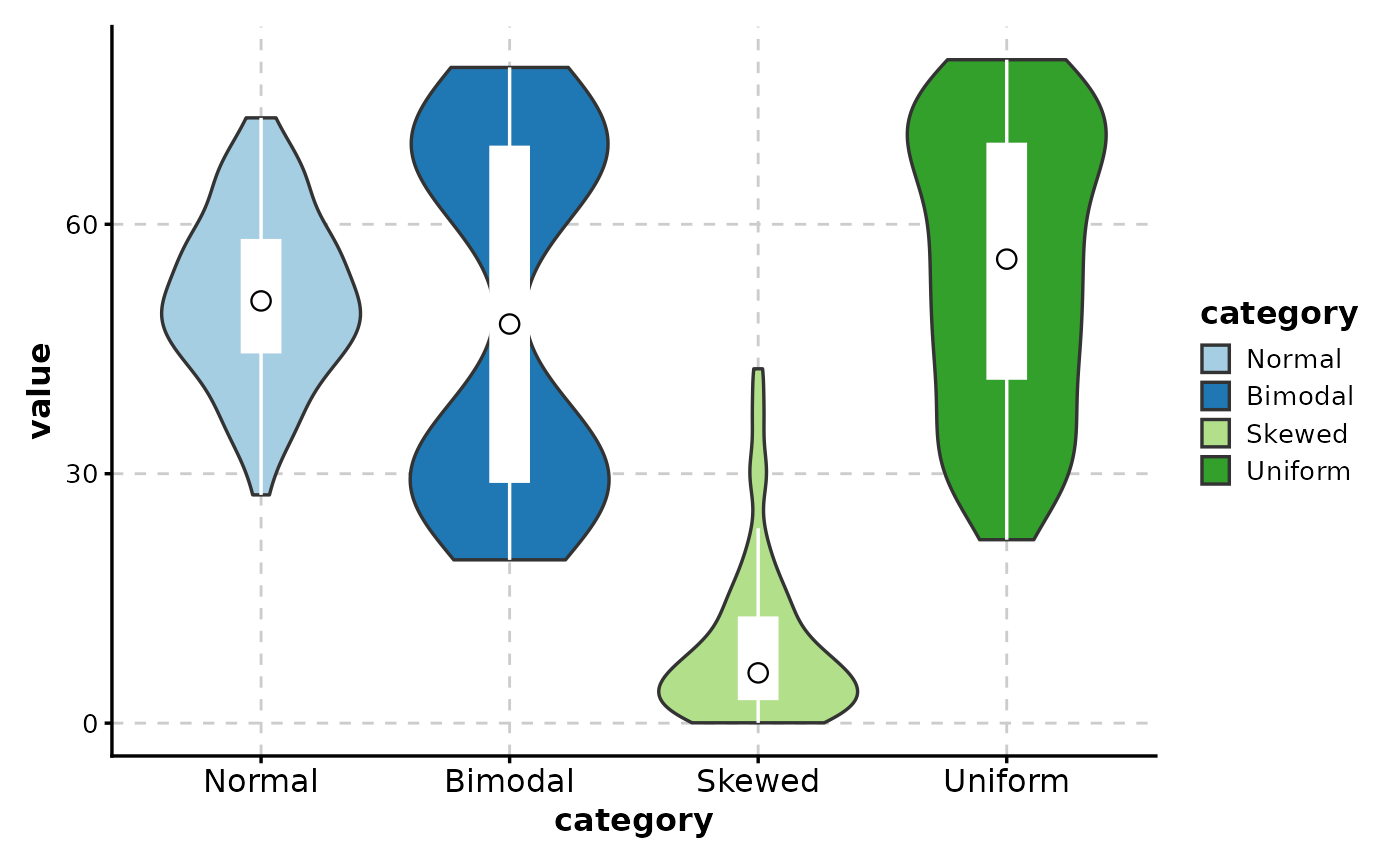
# Customize box overlay
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE,
box_color = "white",
box_width = 0.15,
box_ptsize = 3
)
 # ============================================================
# Adding Data Points
# ============================================================
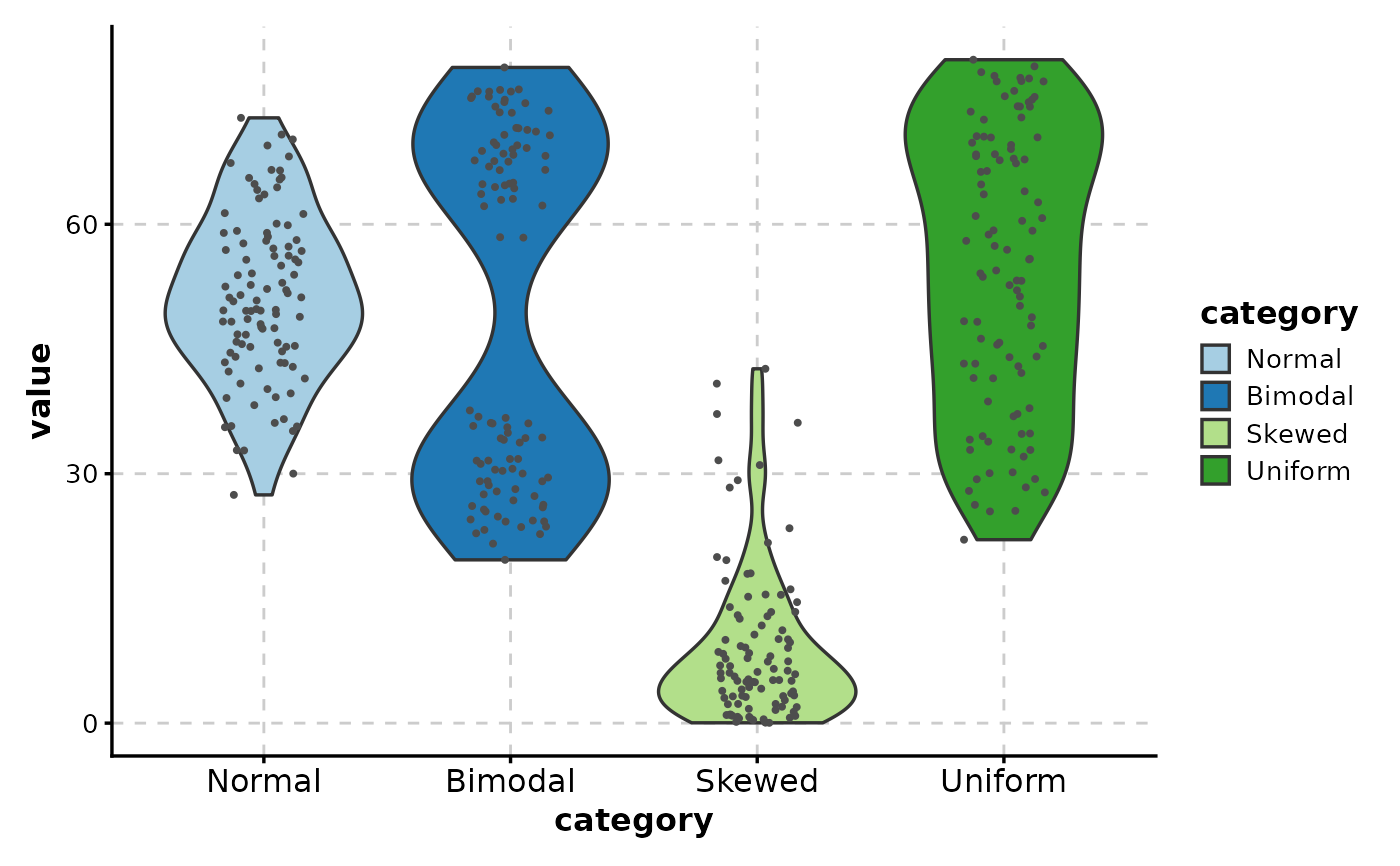
# Add jittered points to show individual observations
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE
)
# ============================================================
# Adding Data Points
# ============================================================
# Add jittered points to show individual observations
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE
)
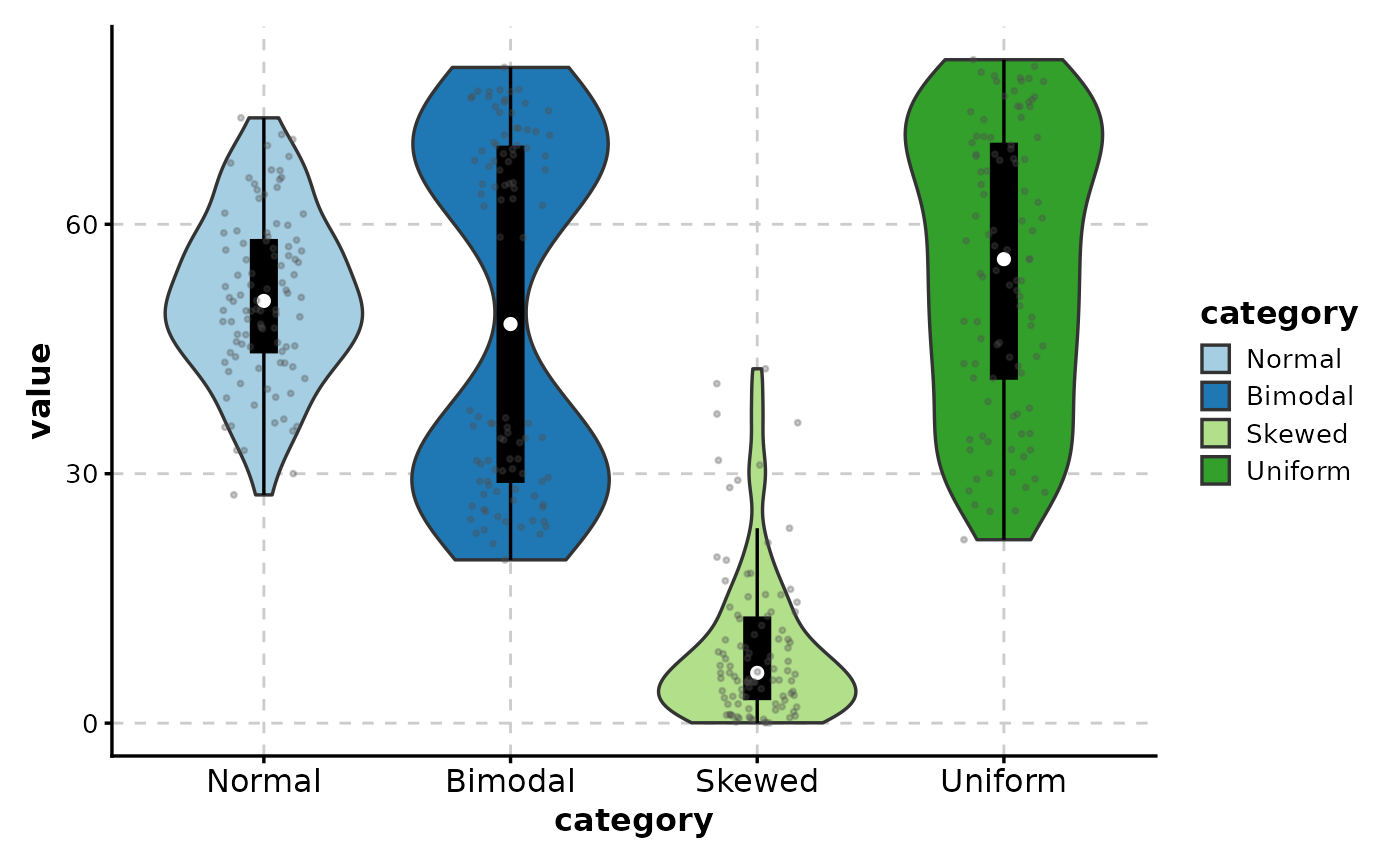
 # Combine box overlay with points
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE,
add_point = TRUE,
pt_alpha = 0.3
)
# Combine box overlay with points
ViolinPlot(data,
x = "category", y = "value",
add_box = TRUE,
add_point = TRUE,
pt_alpha = 0.3
)
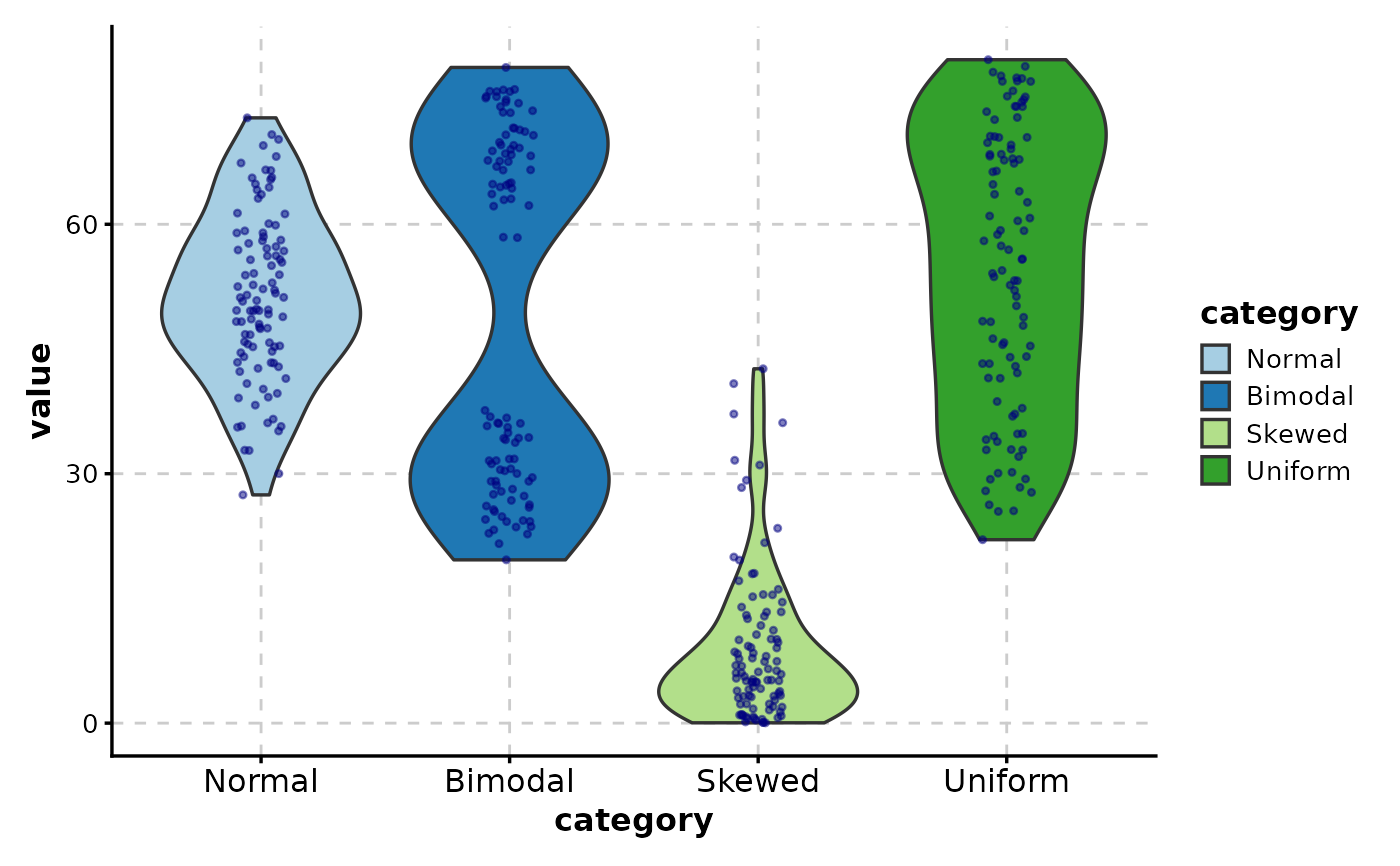
 # Customize point appearance
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE,
pt_color = "navy",
pt_size = 0.8,
pt_alpha = 0.5,
jitter_width = 0.3
)
# Customize point appearance
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE,
pt_color = "navy",
pt_size = 0.8,
pt_alpha = 0.5,
jitter_width = 0.3
)
 # ============================================================
# Grouped Violin Plots
# ============================================================
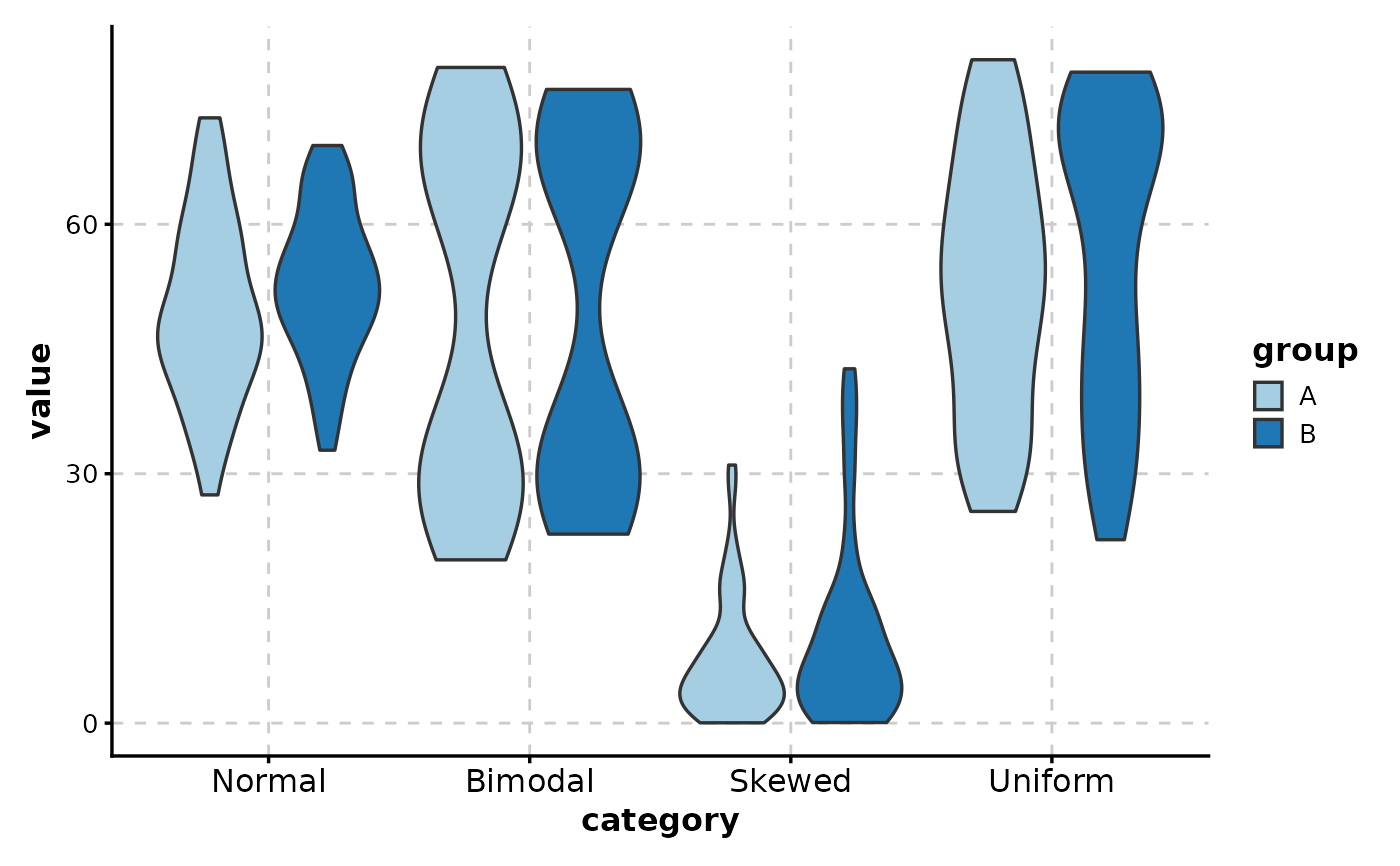
# Side-by-side violins by group
ViolinPlot(data,
x = "category", y = "value",
group_by = "group"
)
# ============================================================
# Grouped Violin Plots
# ============================================================
# Side-by-side violins by group
ViolinPlot(data,
x = "category", y = "value",
group_by = "group"
)
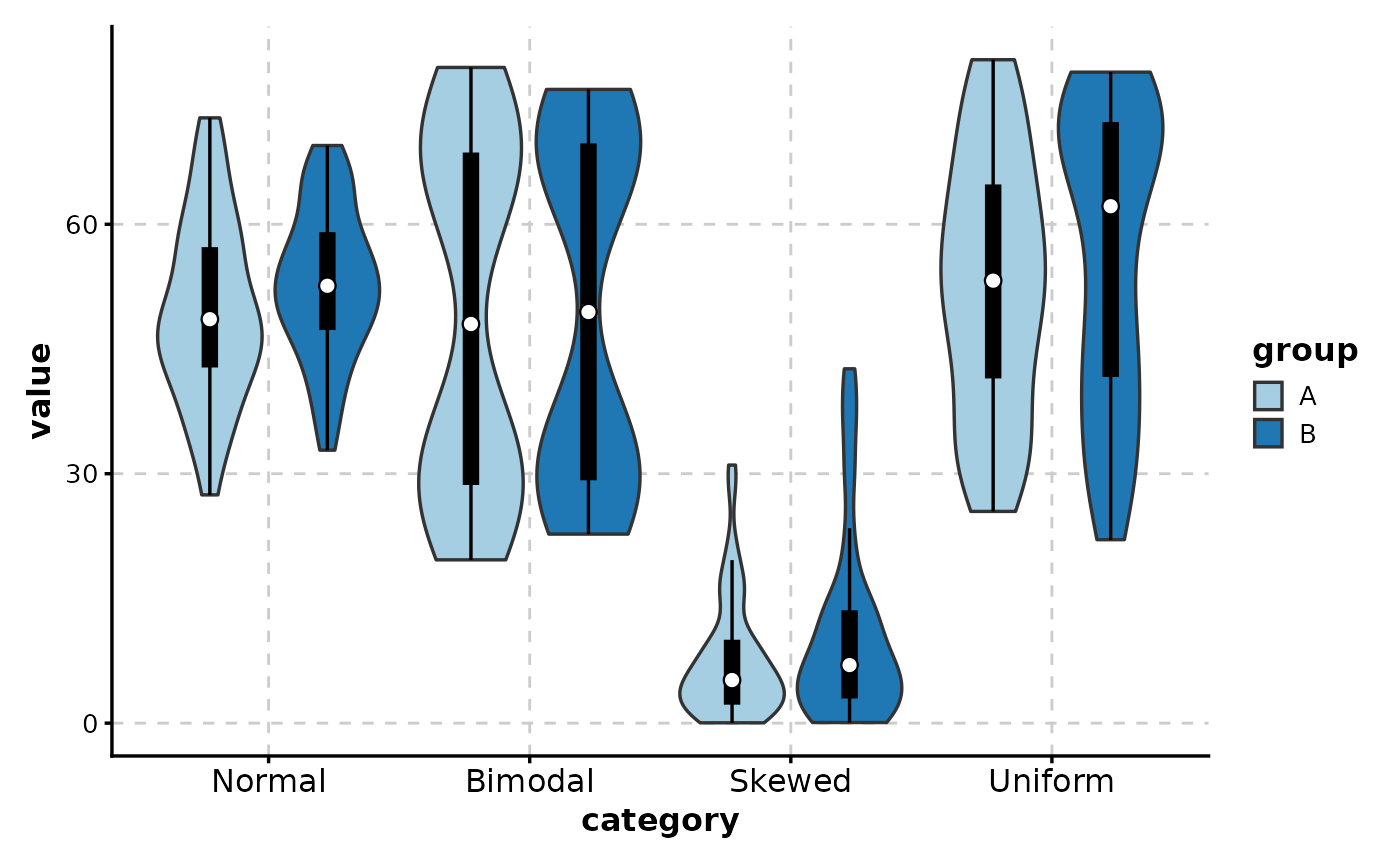
 # With box overlay
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
add_box = TRUE
)
# With box overlay
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
add_box = TRUE
)
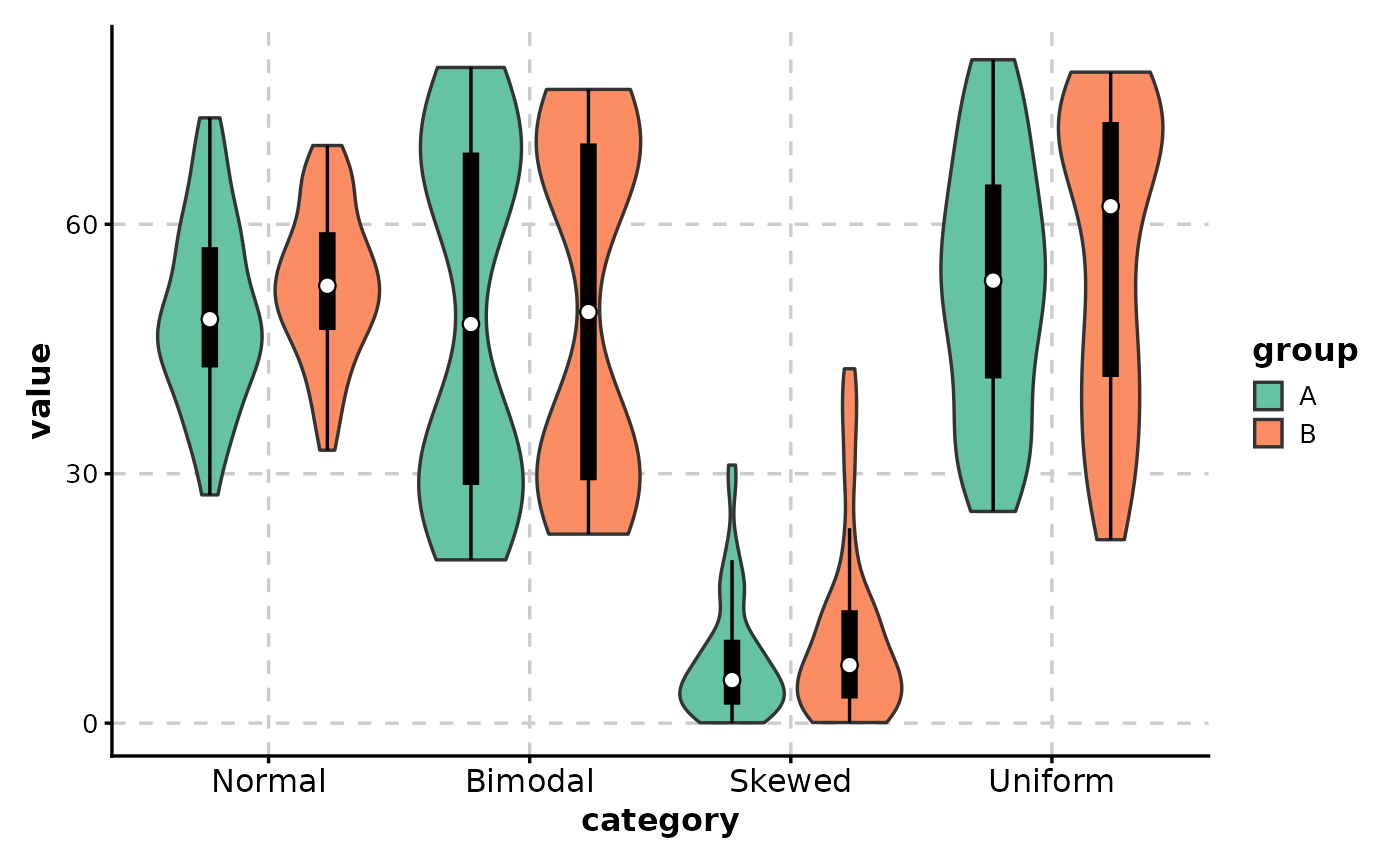
 # Custom palette
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
palette = "Set2",
add_box = TRUE
)
# Custom palette
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
palette = "Set2",
add_box = TRUE
)
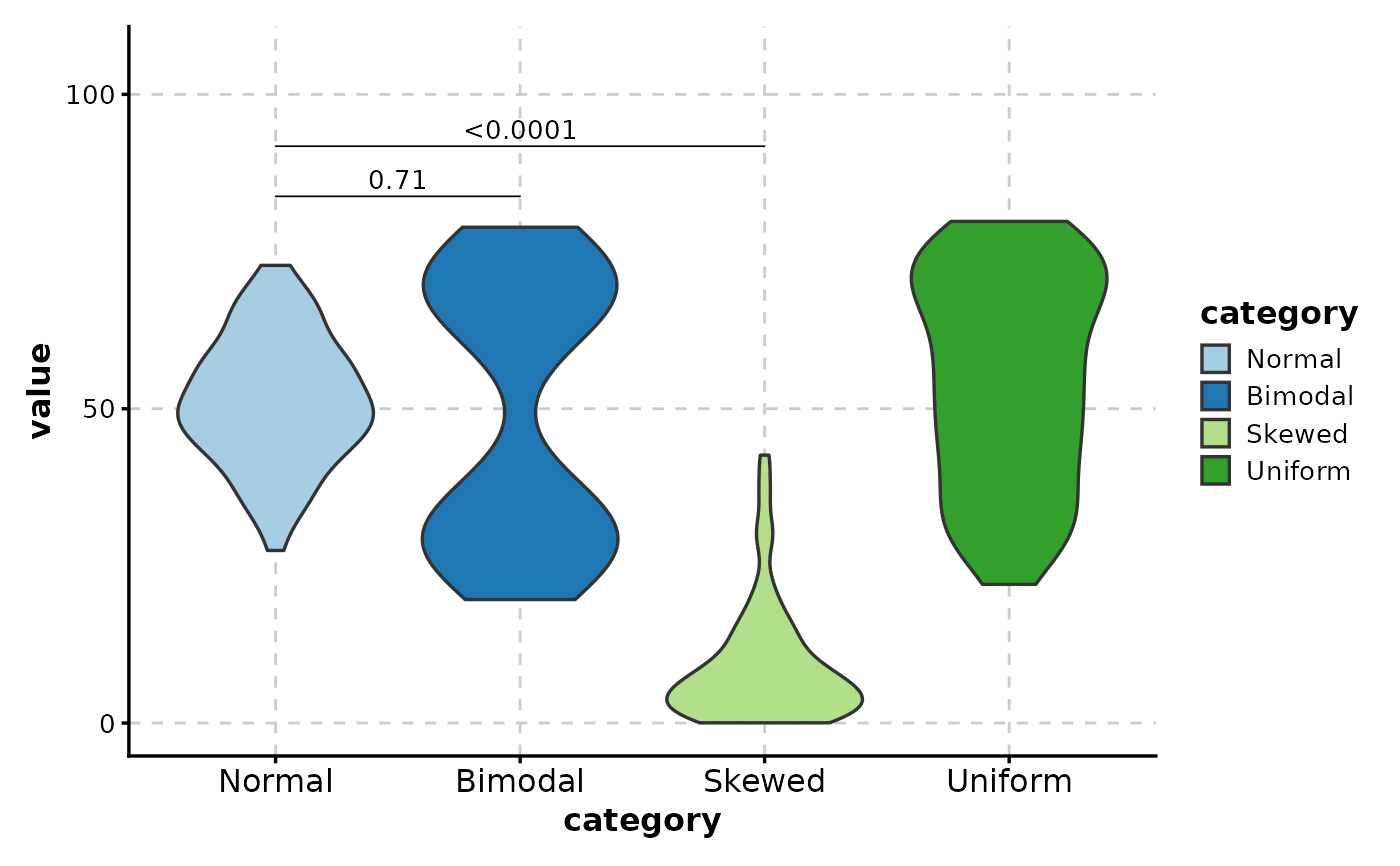
 # ============================================================
# Statistical Comparisons
# ============================================================
# Compare specific distributions
ViolinPlot(data,
x = "category", y = "value",
comparisons = list(
c("Normal", "Bimodal"),
c("Normal", "Skewed")
)
)
# ============================================================
# Statistical Comparisons
# ============================================================
# Compare specific distributions
ViolinPlot(data,
x = "category", y = "value",
comparisons = list(
c("Normal", "Bimodal"),
c("Normal", "Skewed")
)
)
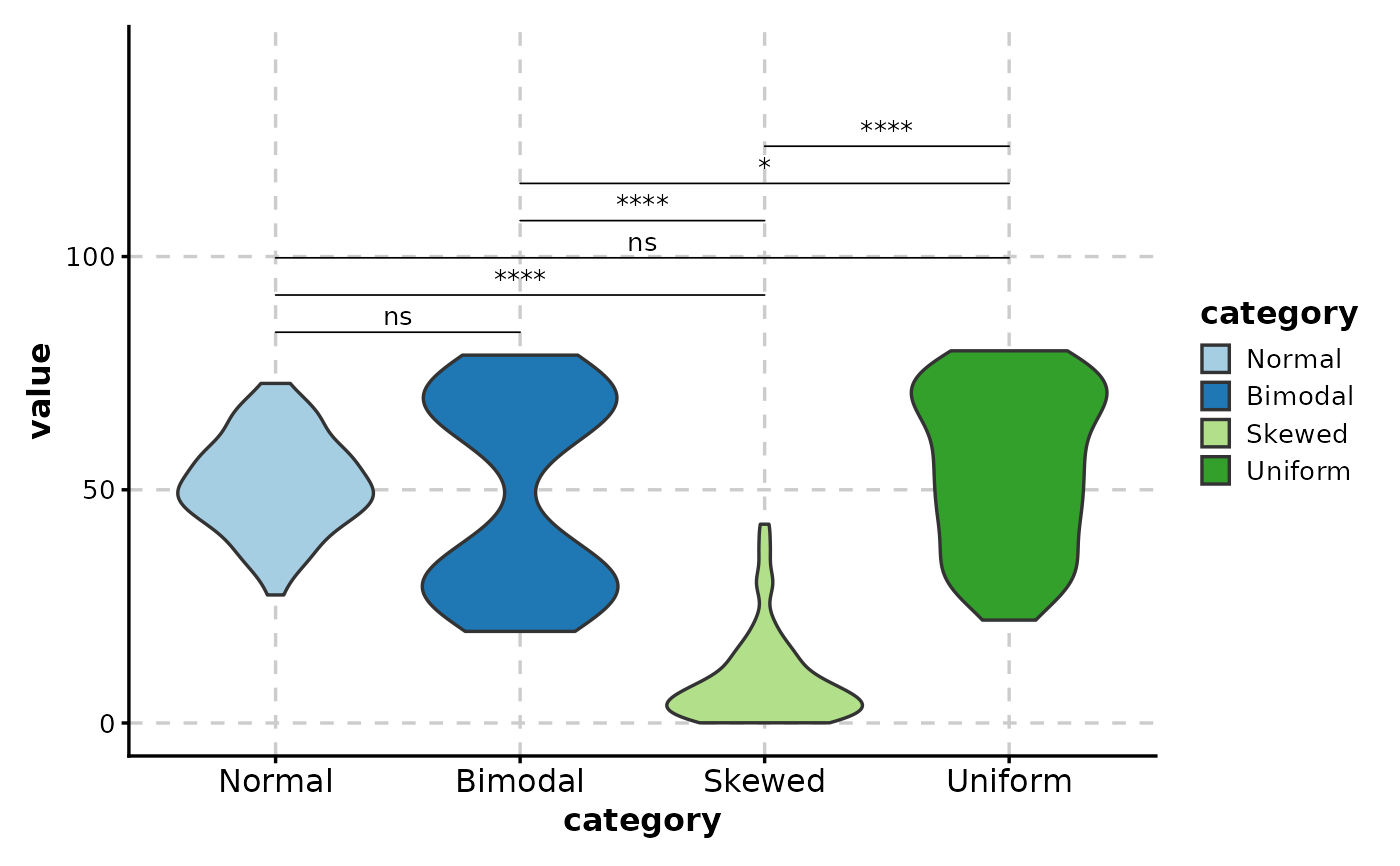
 # All pairwise comparisons
ViolinPlot(data,
x = "category", y = "value",
comparisons = TRUE,
sig_label = "p.signif"
)
# All pairwise comparisons
ViolinPlot(data,
x = "category", y = "value",
comparisons = TRUE,
sig_label = "p.signif"
)
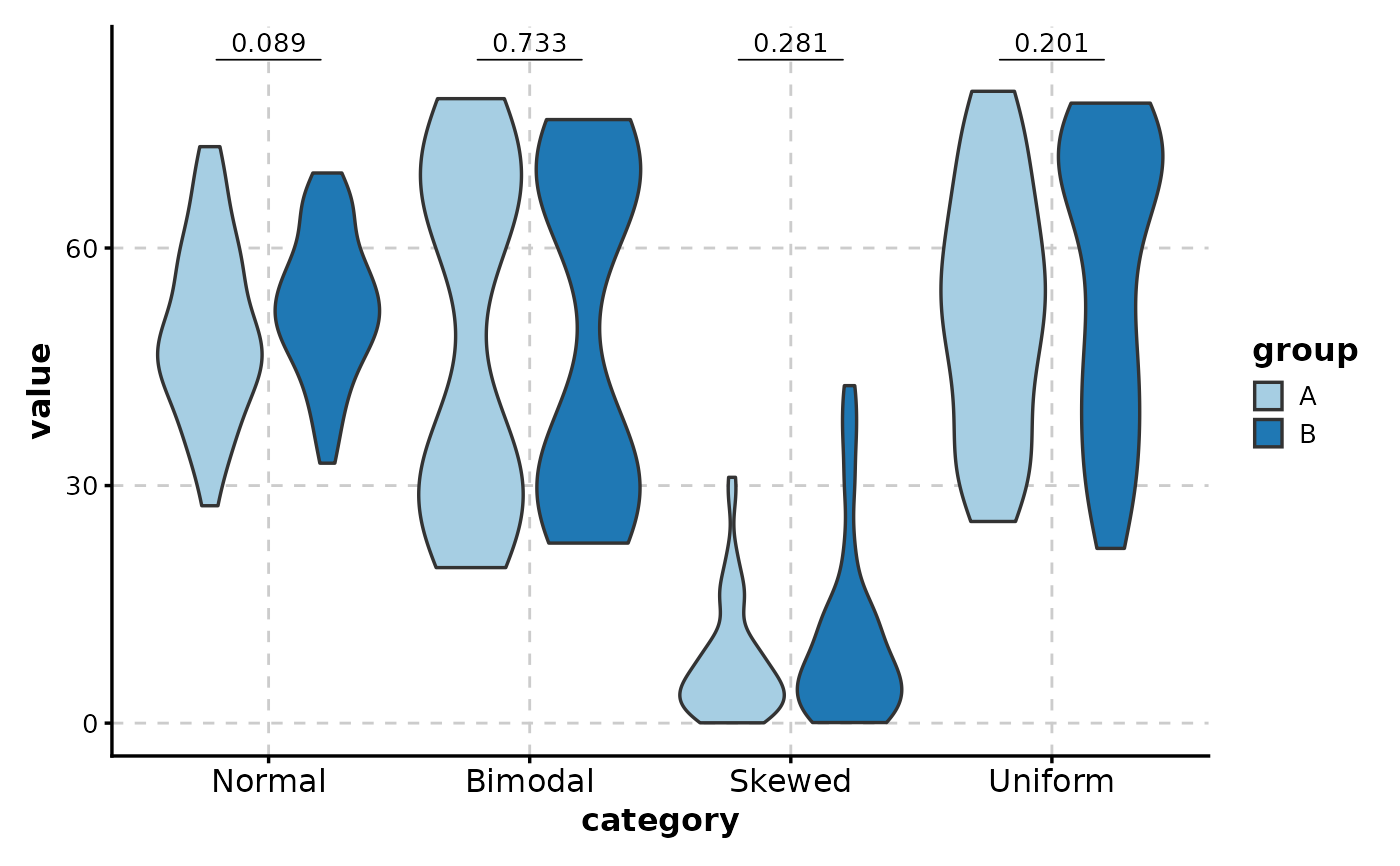
 # Grouped comparisons
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
comparisons = TRUE
)
# Grouped comparisons
ViolinPlot(data,
x = "category", y = "value",
group_by = "group",
comparisons = TRUE
)
 # ============================================================
# Paired Data Analysis
# ============================================================
# Create paired data
paired_data <- data.frame(
condition = factor(rep(c("Baseline", "Treatment"), each = 30)),
patient = factor(rep(1:30, 2)),
response = c(rnorm(30, 100, 20), rnorm(30, 120, 20))
)
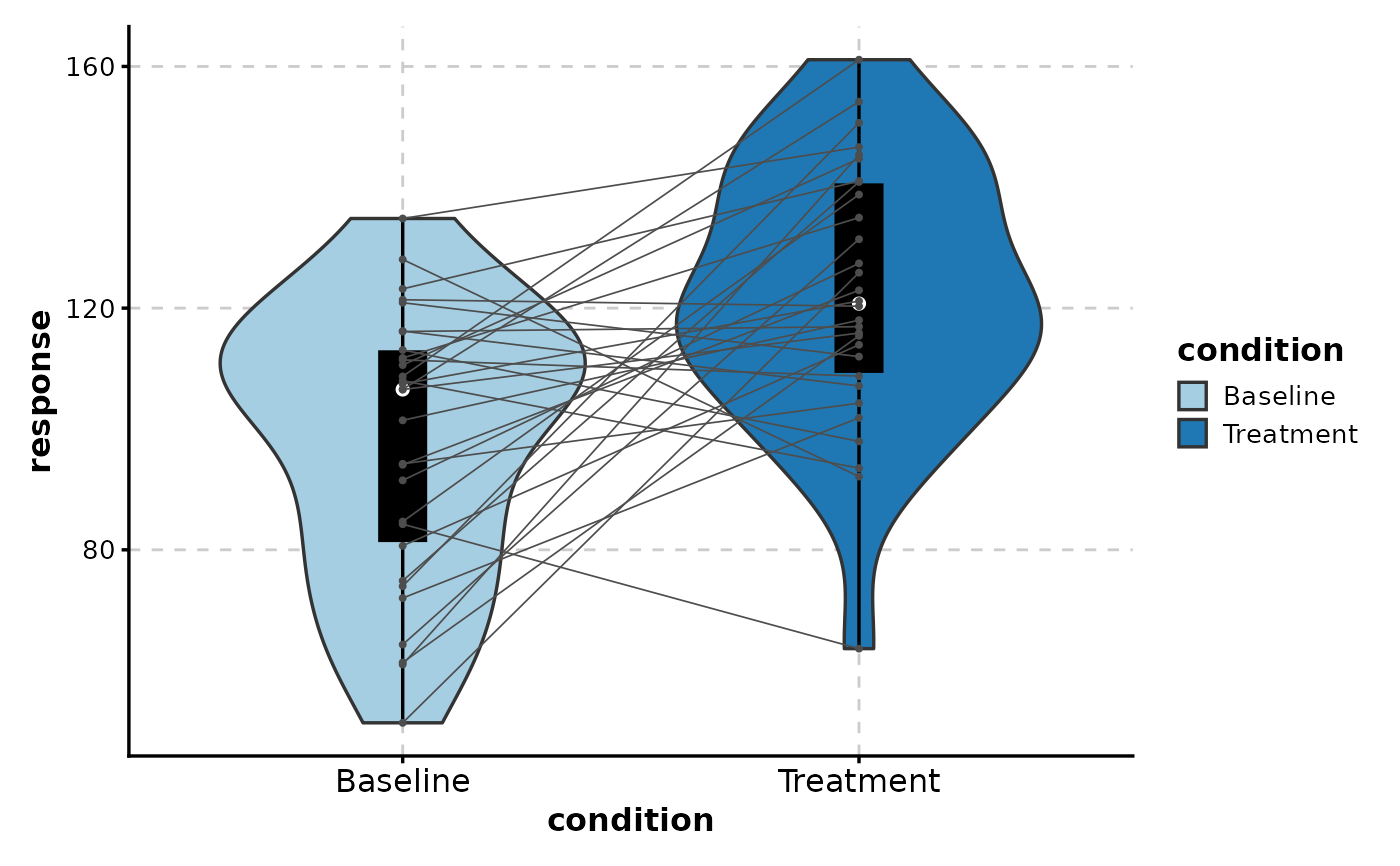
# Paired violin with connecting lines
ViolinPlot(paired_data,
x = "condition", y = "response",
paired_by = "patient",
add_box = TRUE
)
#> Warning: Forcing `add_point = TRUE` when `paired_by` is provided.
# ============================================================
# Paired Data Analysis
# ============================================================
# Create paired data
paired_data <- data.frame(
condition = factor(rep(c("Baseline", "Treatment"), each = 30)),
patient = factor(rep(1:30, 2)),
response = c(rnorm(30, 100, 20), rnorm(30, 120, 20))
)
# Paired violin with connecting lines
ViolinPlot(paired_data,
x = "condition", y = "response",
paired_by = "patient",
add_box = TRUE
)
#> Warning: Forcing `add_point = TRUE` when `paired_by` is provided.
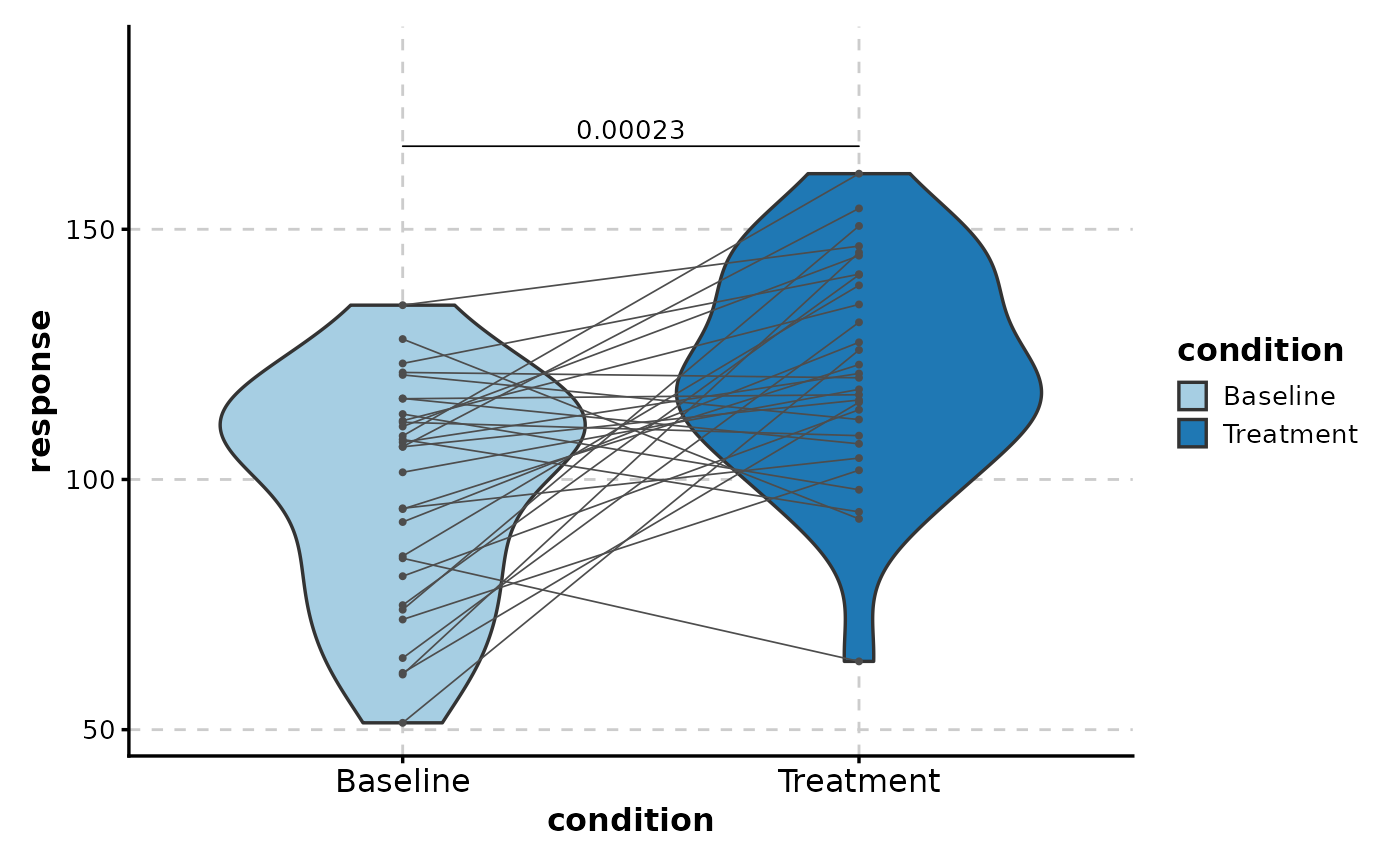
 # With paired t-test
ViolinPlot(paired_data,
x = "condition", y = "response",
paired_by = "patient",
comparisons = list(c("Baseline", "Treatment")),
pairwise_method = "t.test"
)
#> Warning: Forcing `add_point = TRUE` when `paired_by` is provided.
# With paired t-test
ViolinPlot(paired_data,
x = "condition", y = "response",
paired_by = "patient",
comparisons = list(c("Baseline", "Treatment")),
pairwise_method = "t.test"
)
#> Warning: Forcing `add_point = TRUE` when `paired_by` is provided.
 # ============================================================
# Highlighting and Visual Enhancements
# ============================================================
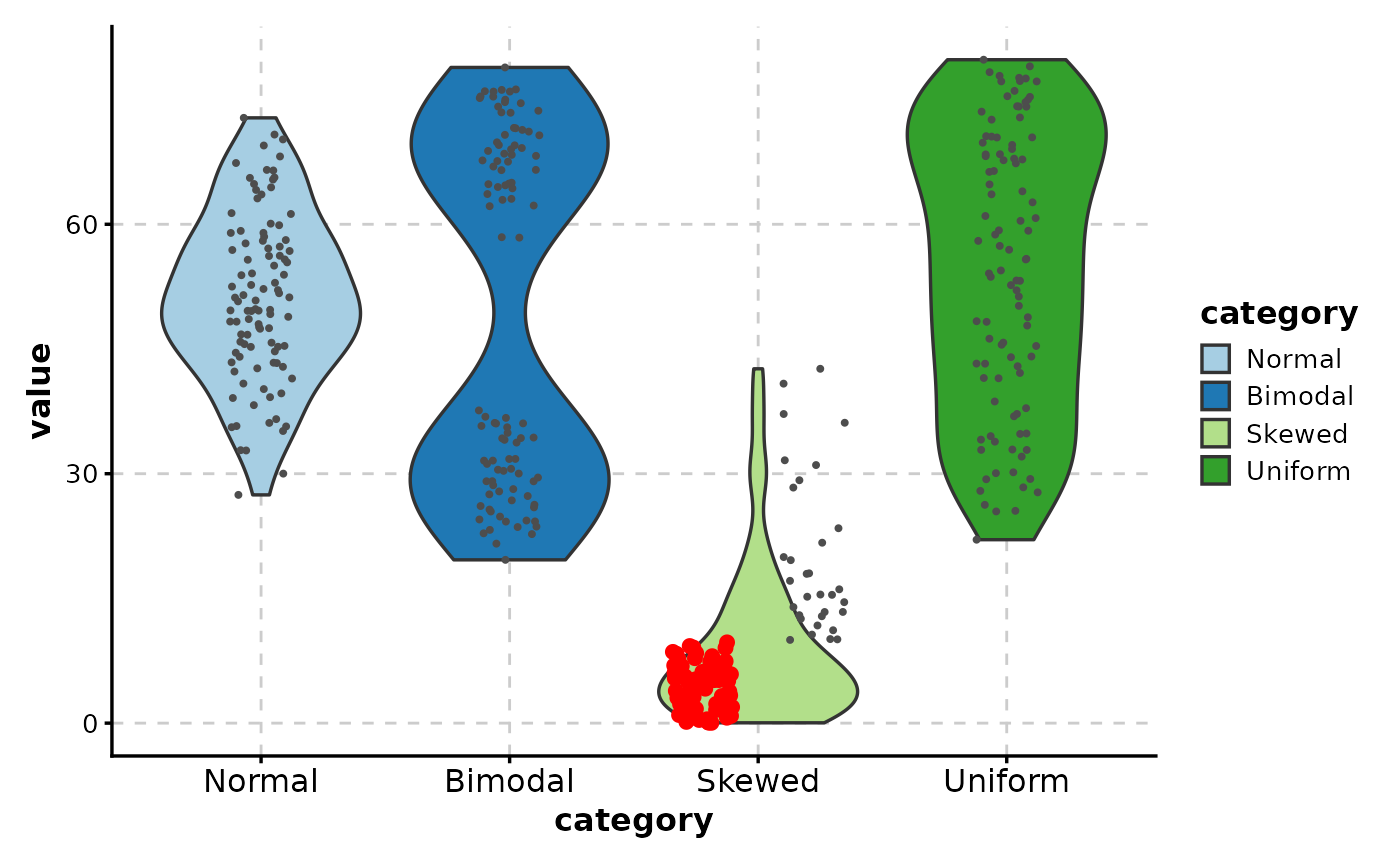
# Highlight extreme values
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE,
highlight = "value > 80 | value < 10",
highlight_color = "red",
highlight_size = 2
)
# ============================================================
# Highlighting and Visual Enhancements
# ============================================================
# Highlight extreme values
ViolinPlot(data,
x = "category", y = "value",
add_point = TRUE,
highlight = "value > 80 | value < 10",
highlight_color = "red",
highlight_size = 2
)
 # Add trend line
ViolinPlot(data,
x = "category", y = "value",
add_trend = TRUE,
trend_linewidth = 1.5
)
# Add trend line
ViolinPlot(data,
x = "category", y = "value",
add_trend = TRUE,
trend_linewidth = 1.5
)
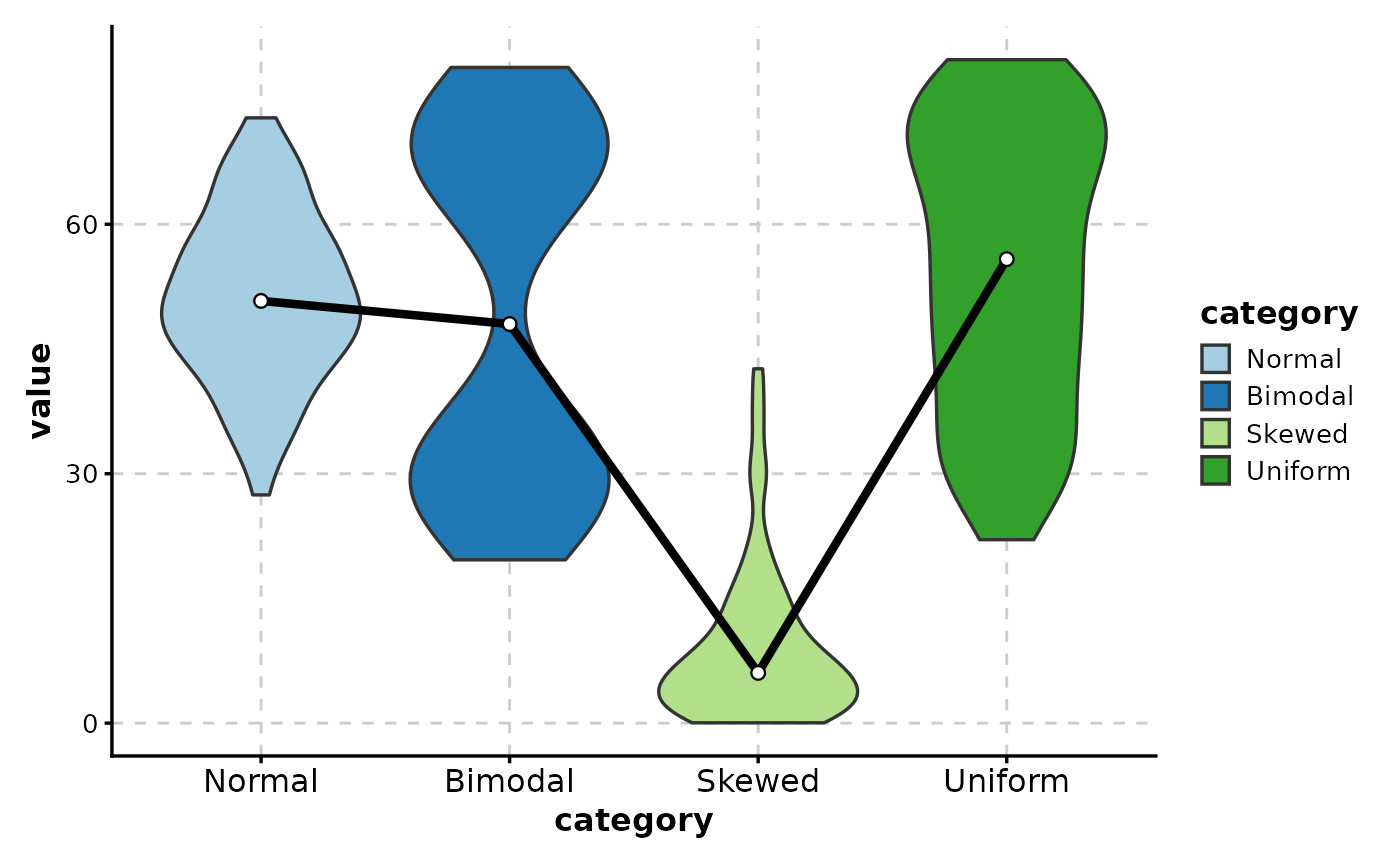 # Add reference line
ViolinPlot(data,
x = "category", y = "value",
add_line = 50,
line_color = "darkgreen",
line_type = 1
)
# Add reference line
ViolinPlot(data,
x = "category", y = "value",
add_line = 50,
line_color = "darkgreen",
line_type = 1
)
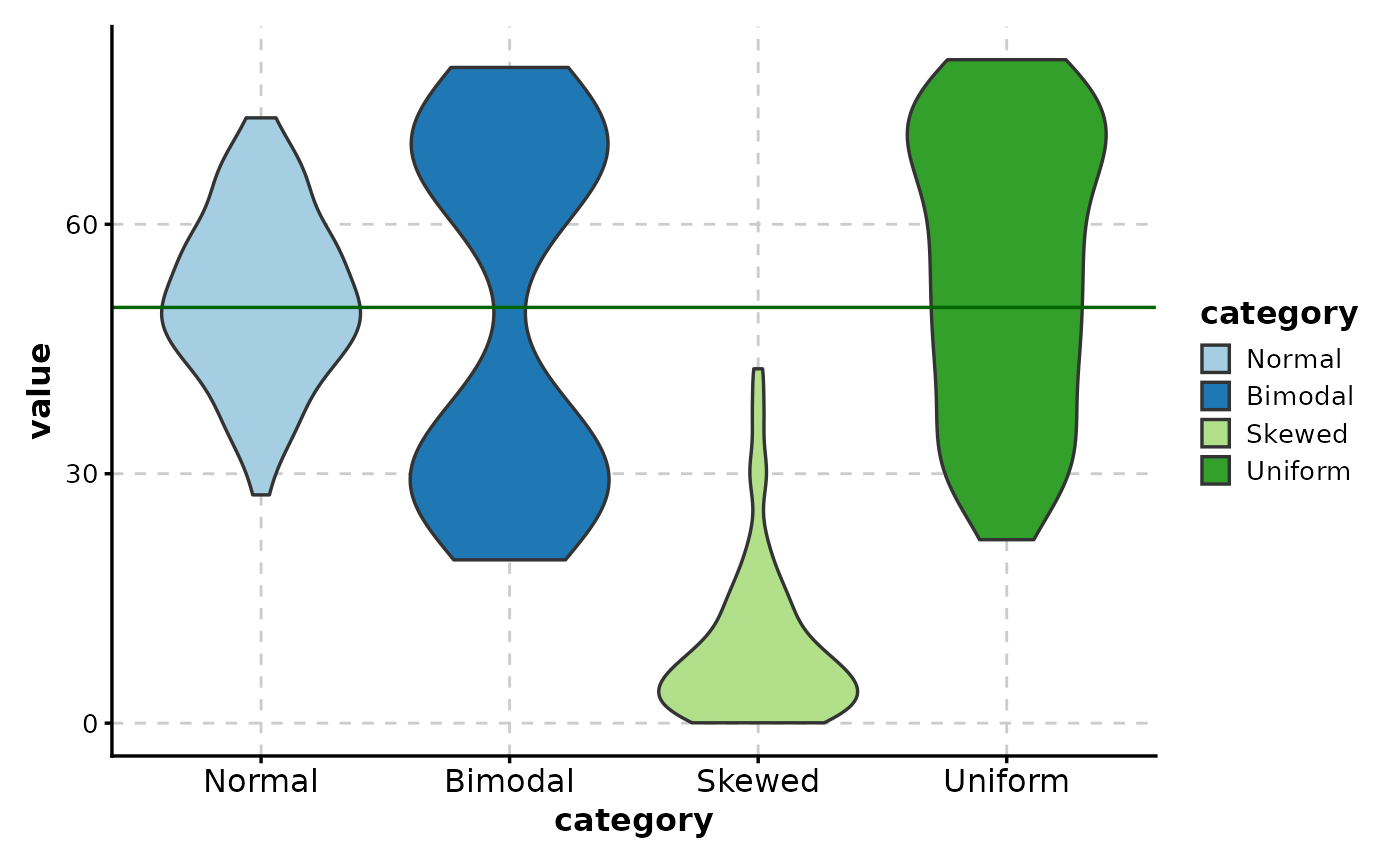 # Add mean indicator
ViolinPlot(data,
x = "category", y = "value",
add_stat = mean,
stat_name = "Mean",
stat_color = "red",
stat_shape = 18,
stat_size = 3
)
# Add mean indicator
ViolinPlot(data,
x = "category", y = "value",
add_stat = mean,
stat_name = "Mean",
stat_color = "red",
stat_shape = 18,
stat_size = 3
)
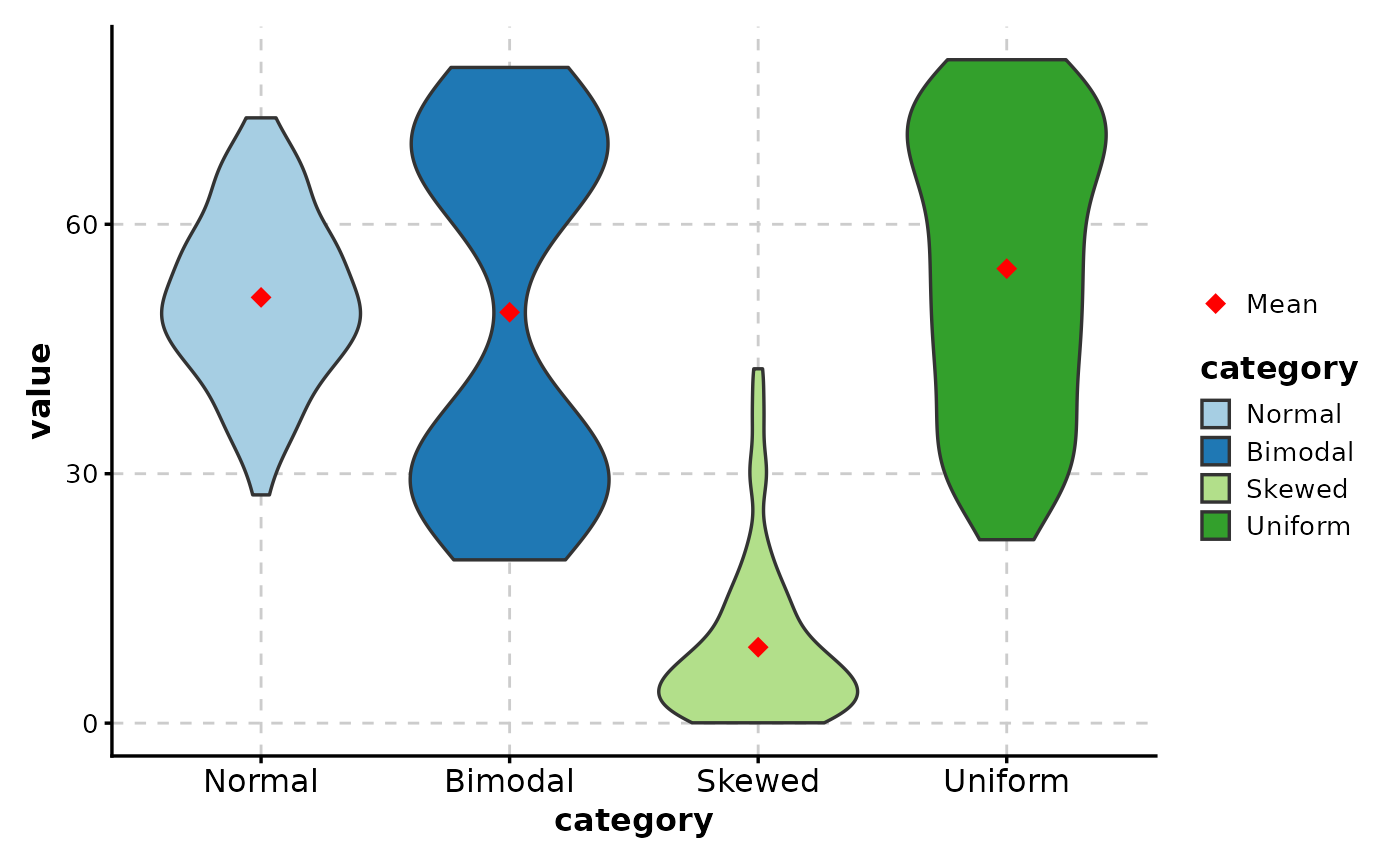 # ============================================================
# Fill Modes
# ============================================================
# Fill by x category
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "x",
palette = "Pastel1"
)
# ============================================================
# Fill Modes
# ============================================================
# Fill by x category
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "x",
palette = "Pastel1"
)
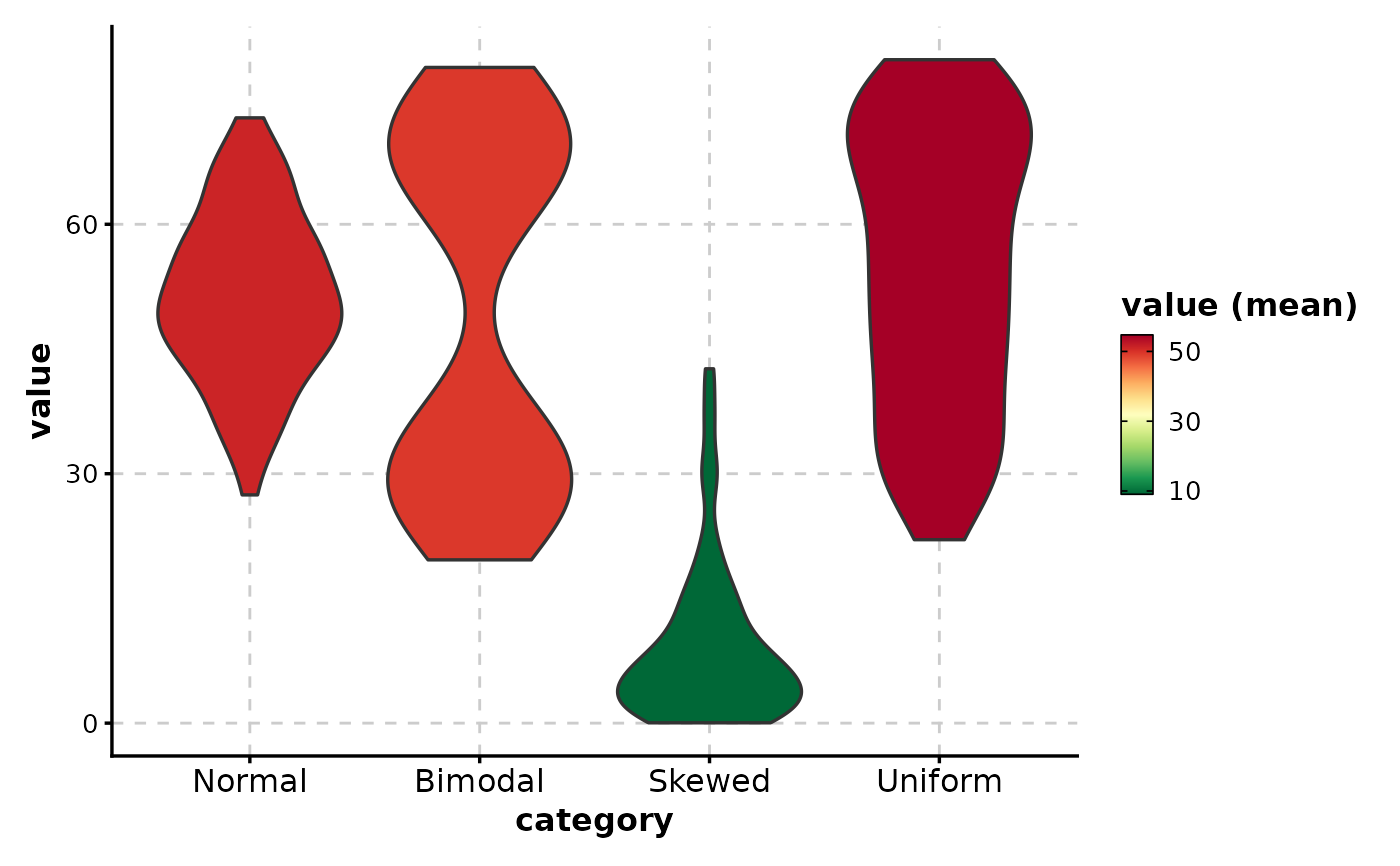
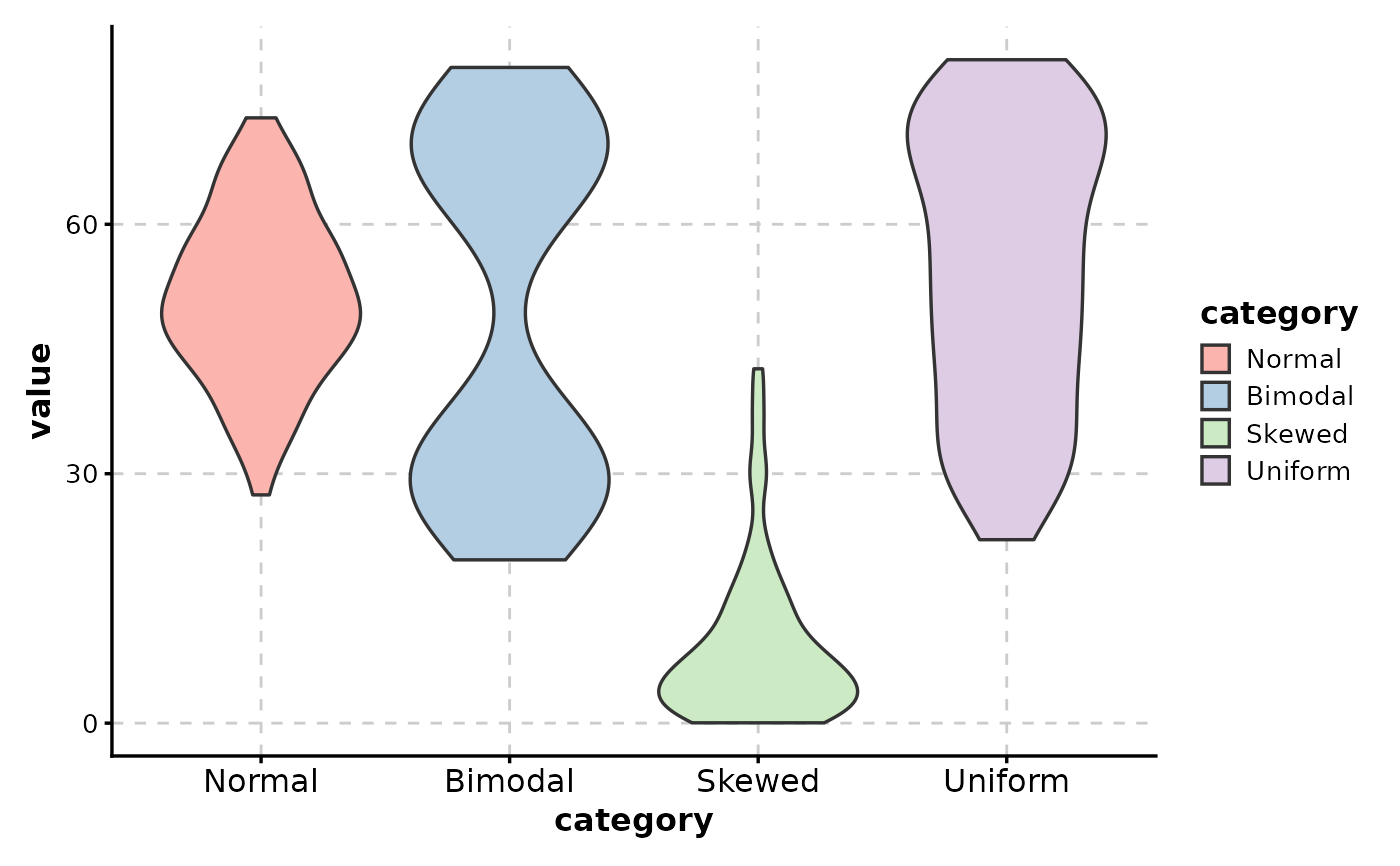 # Fill by mean (gradient coloring)
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "mean",
palette = "RdYlGn"
)
# Fill by mean (gradient coloring)
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "mean",
palette = "RdYlGn"
)
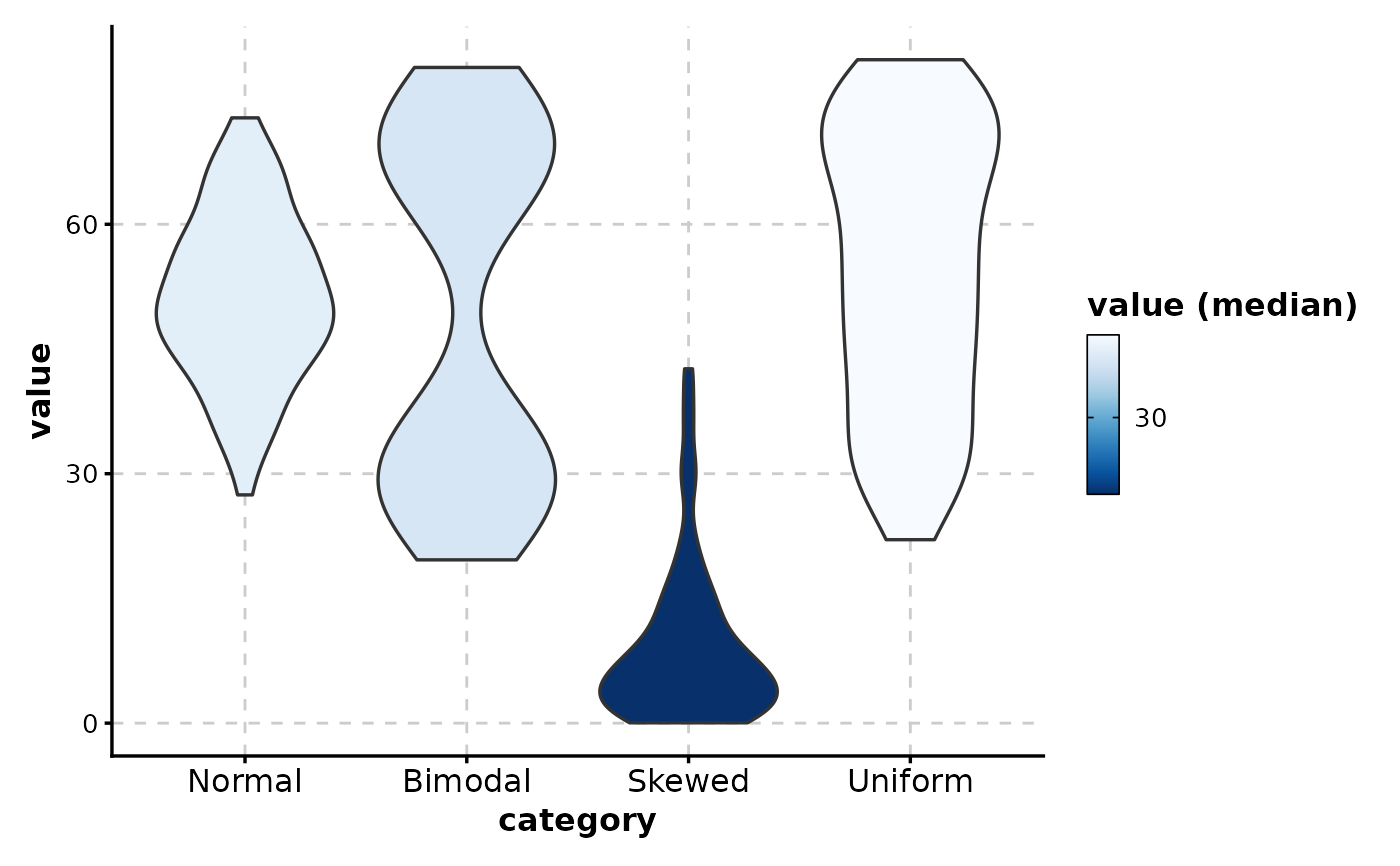
 # Fill by median with reversed gradient
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "median",
palette = "Blues",
fill_reverse = TRUE
)
# Fill by median with reversed gradient
ViolinPlot(data,
x = "category", y = "value",
fill_mode = "median",
palette = "Blues",
fill_reverse = TRUE
)
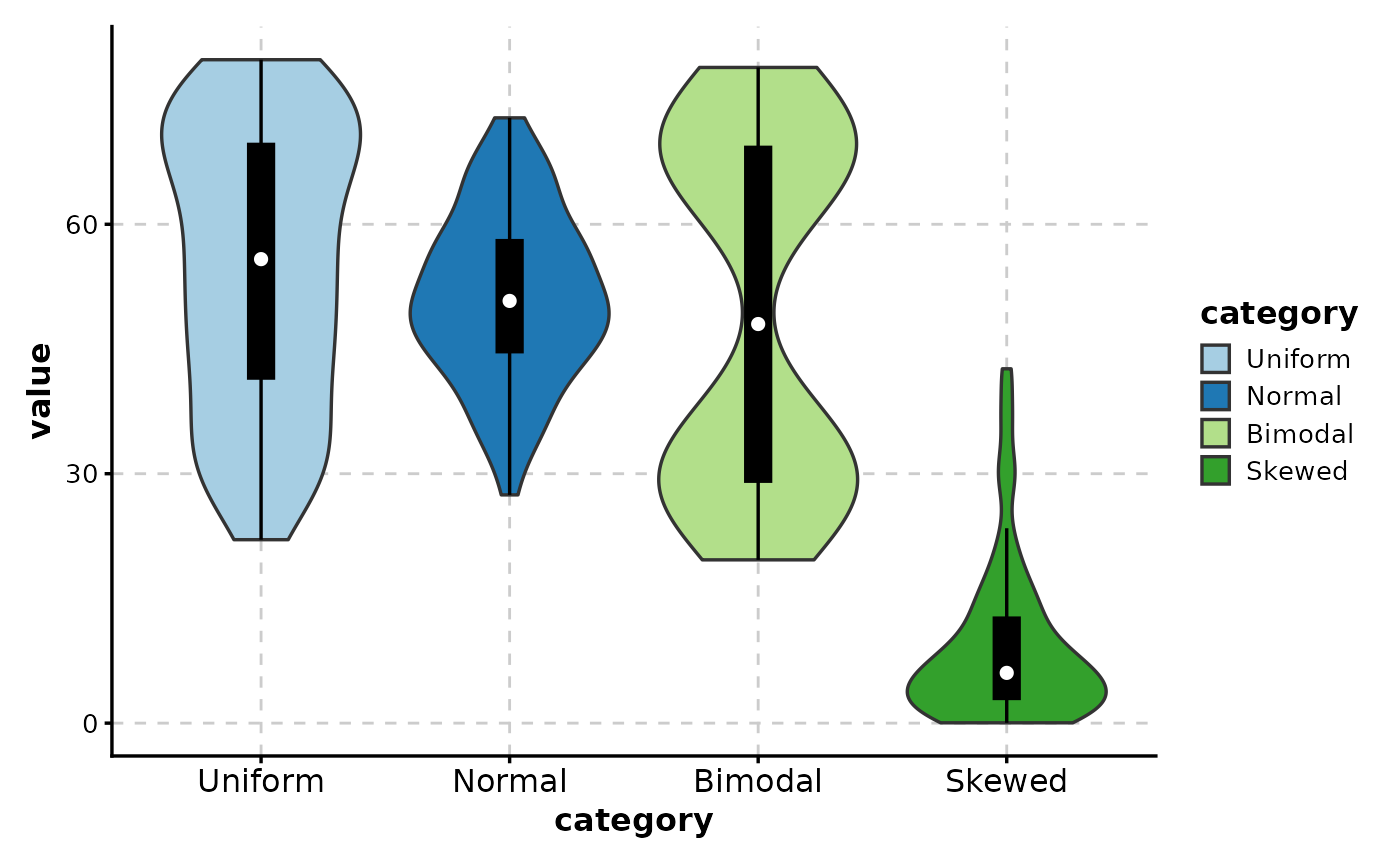
 # ============================================================
# Sorting and Orientation
# ============================================================
# Sort by mean value
ViolinPlot(data,
x = "category", y = "value",
sort_x = "mean_desc",
add_box = TRUE
)
# ============================================================
# Sorting and Orientation
# ============================================================
# Sort by mean value
ViolinPlot(data,
x = "category", y = "value",
sort_x = "mean_desc",
add_box = TRUE
)
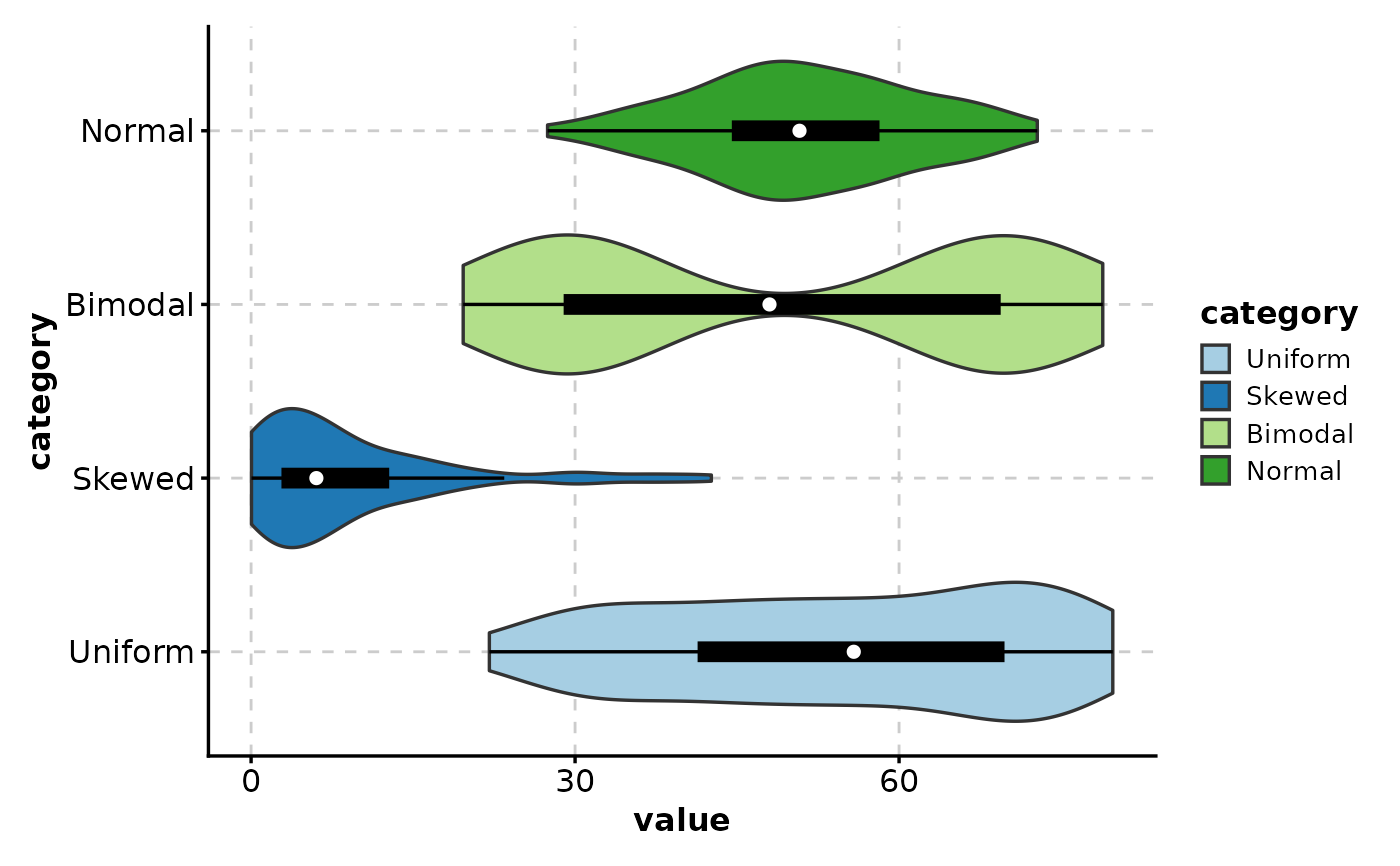
 # Horizontal violin plot
ViolinPlot(data,
x = "category", y = "value",
flip = TRUE,
add_box = TRUE
)
# Horizontal violin plot
ViolinPlot(data,
x = "category", y = "value",
flip = TRUE,
add_box = TRUE
)
 # ============================================================
# Faceting
# ============================================================
# Add faceting variable
data$experiment <- sample(c("Exp1", "Exp2"), nrow(data), replace = TRUE)
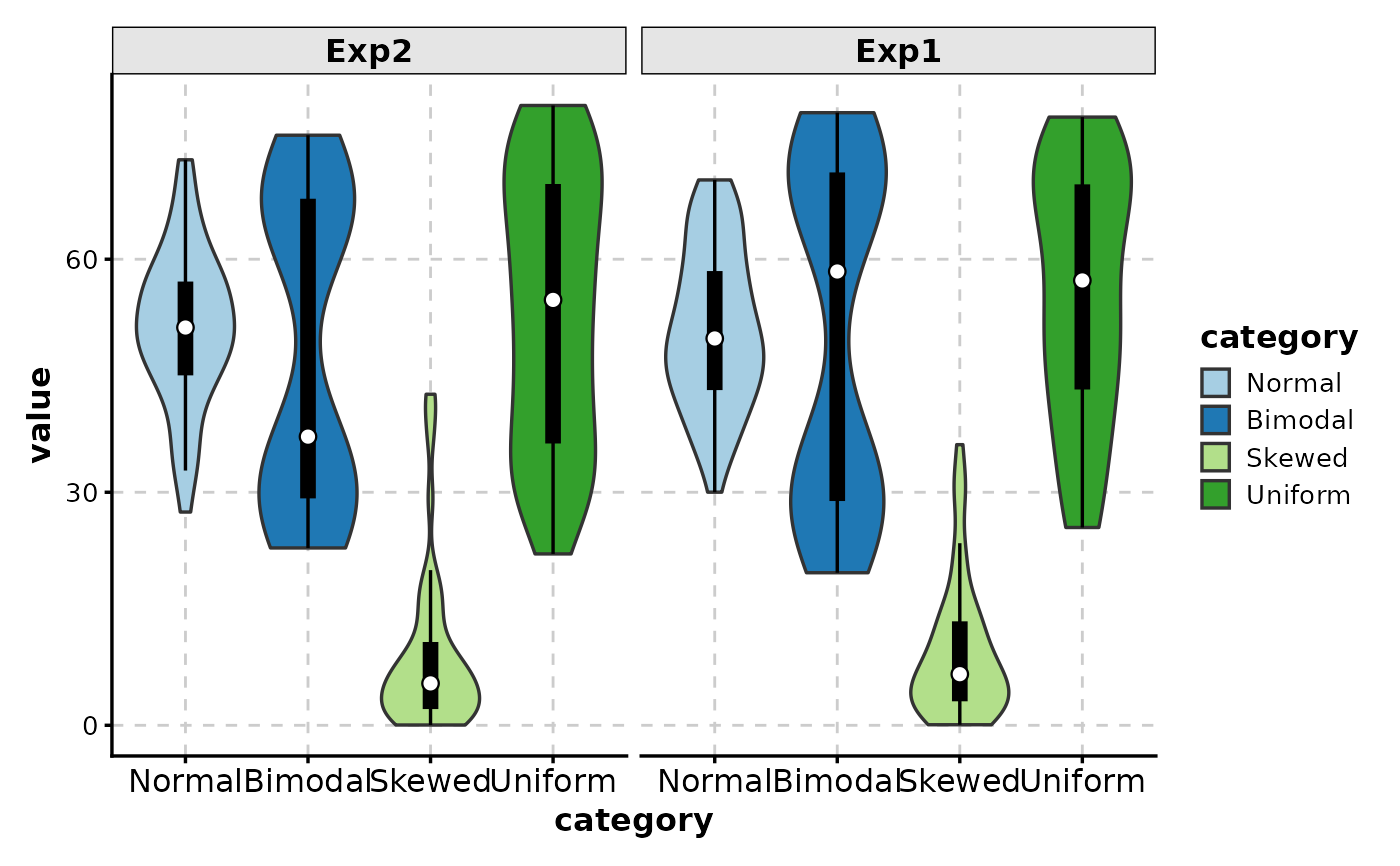
# Facet by experiment
ViolinPlot(data,
x = "category", y = "value",
facet_by = "experiment",
add_box = TRUE
)
# ============================================================
# Faceting
# ============================================================
# Add faceting variable
data$experiment <- sample(c("Exp1", "Exp2"), nrow(data), replace = TRUE)
# Facet by experiment
ViolinPlot(data,
x = "category", y = "value",
facet_by = "experiment",
add_box = TRUE
)
 # Free scales
ViolinPlot(data,
x = "category", y = "value",
facet_by = "experiment",
facet_scales = "free_y"
)
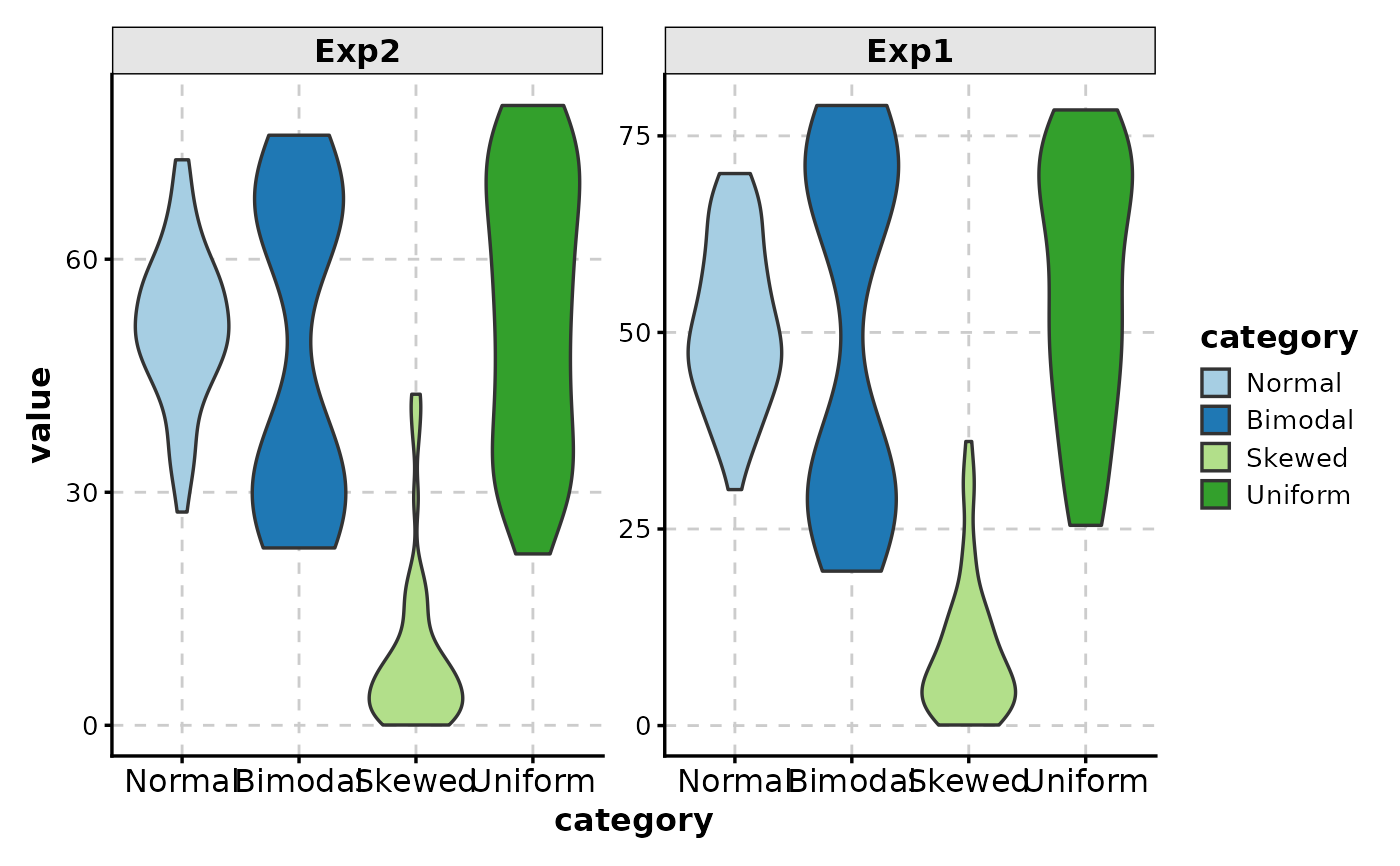
# Free scales
ViolinPlot(data,
x = "category", y = "value",
facet_by = "experiment",
facet_scales = "free_y"
)
 # ============================================================
# Wide Format Data
# ============================================================
# Wide format input
wide_data <- data.frame(
Control = rnorm(50, 100, 15),
LowDose = rnorm(50, 110, 15),
HighDose = rnorm(50, 130, 20)
)
ViolinPlot(wide_data,
x = c("Control", "LowDose", "HighDose"),
in_form = "wide",
add_box = TRUE,
xlab = "Treatment",
ylab = "Response"
)
# ============================================================
# Wide Format Data
# ============================================================
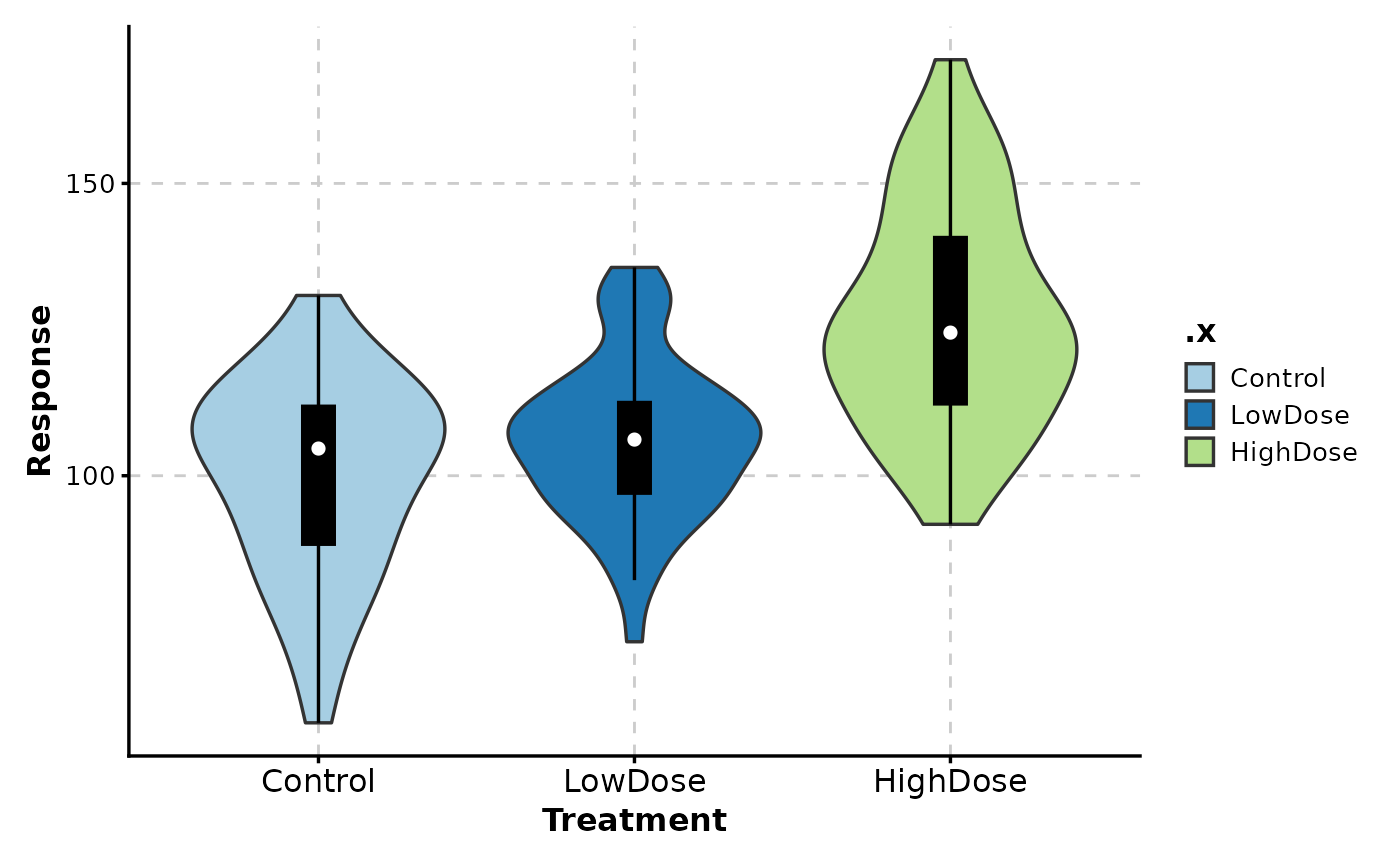
# Wide format input
wide_data <- data.frame(
Control = rnorm(50, 100, 15),
LowDose = rnorm(50, 110, 15),
HighDose = rnorm(50, 130, 20)
)
ViolinPlot(wide_data,
x = c("Control", "LowDose", "HighDose"),
in_form = "wide",
add_box = TRUE,
xlab = "Treatment",
ylab = "Response"
)
 # ============================================================
# Complex Example: Publication-Ready Plot
# ============================================================
# Gene expression data example
expr_data <- data.frame(
gene = rep(c("BRCA1", "TP53", "EGFR", "MYC"), each = 40),
expression = c(
rnorm(40, 8, 1.5),
rnorm(40, 6, 2),
rnorm(40, 10, 1),
rnorm(40, 12, 2.5)
),
tissue = rep(rep(c("Normal", "Tumor"), each = 20), 4)
)
ViolinPlot(expr_data,
x = "gene",
y = "expression",
group_by = "tissue",
add_box = TRUE,
add_point = TRUE,
pt_alpha = 0.4,
comparisons = TRUE,
sig_label = "p.signif",
hide_ns = TRUE,
palette = c("Normal" = "#4DAF4A", "Tumor" = "#E41A1C"),
title = "Gene Expression by Tissue Type",
xlab = "Gene",
ylab = "Expression (log2)",
legend.position = "bottom"
)
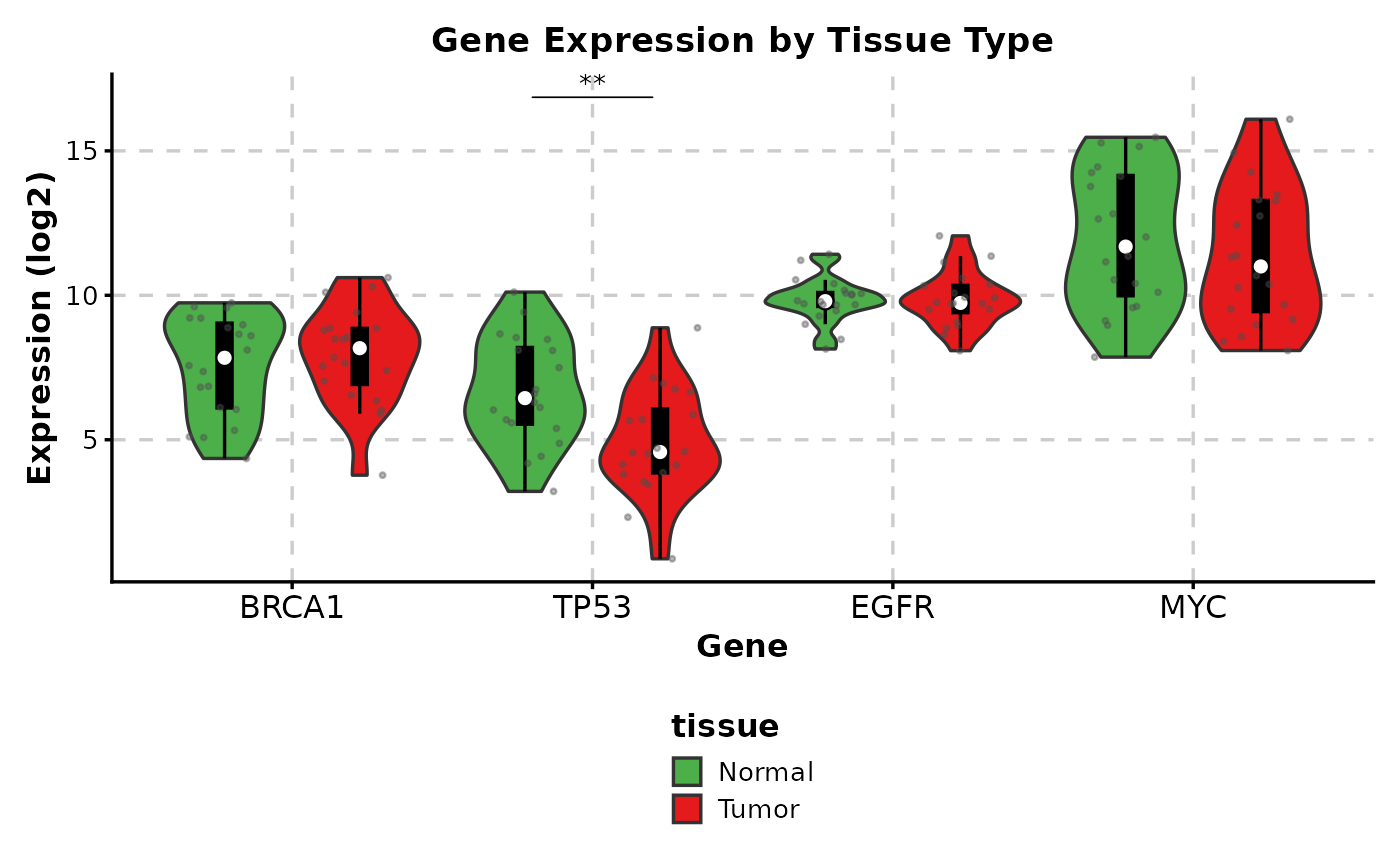
# ============================================================
# Complex Example: Publication-Ready Plot
# ============================================================
# Gene expression data example
expr_data <- data.frame(
gene = rep(c("BRCA1", "TP53", "EGFR", "MYC"), each = 40),
expression = c(
rnorm(40, 8, 1.5),
rnorm(40, 6, 2),
rnorm(40, 10, 1),
rnorm(40, 12, 2.5)
),
tissue = rep(rep(c("Normal", "Tumor"), each = 20), 4)
)
ViolinPlot(expr_data,
x = "gene",
y = "expression",
group_by = "tissue",
add_box = TRUE,
add_point = TRUE,
pt_alpha = 0.4,
comparisons = TRUE,
sig_label = "p.signif",
hide_ns = TRUE,
palette = c("Normal" = "#4DAF4A", "Tumor" = "#E41A1C"),
title = "Gene Expression by Tissue Type",
xlab = "Gene",
ylab = "Expression (log2)",
legend.position = "bottom"
)
 # }
# }
